 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G101499_P01 | ||||||||
| Common Name | Zm.237 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 434aa MW: 46035.7 Da PI: 9.0363 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130.5 | 6.2e-41 | 192 | 268 | 1 | 77 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S-- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkq 77
+Cq+egC+adls ak+yhrrhkvCe h+ka+vv++ g++qrfCqqCsrfh lsefDe krsCr+rLa+hn+rrrk+
GRMZM2G101499_P01 192 RCQAEGCKADLSGAKHYHRRHKVCEYHAKASVVATGGKQQRFCQQCSRFHVLSEFDEVKRSCRKRLAEHNRRRRKPP 268
6*************************************************************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 8.2E-34 | 188 | 254 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.359 | 190 | 267 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.96E-38 | 191 | 270 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 7.1E-31 | 193 | 266 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009554 | Biological Process | megasporogenesis | ||||
| GO:0009556 | Biological Process | microsporogenesis | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 434 aa Download sequence Send to blast |
MMTGRLNVAT SSSPGGDDFL FMPMQQHQPP PPPPPPYVGF EHGHGVAGGG SQRVAGGVQH 60 HLYDGLDFAA SALQFQQAEA PHLHQQQLLT LPSSLGPMAP PPPPPPLPMP MPVPGMPGDH 120 VYPALGMVKR EGGGADGRIG LNLGRRTYFS PGDMMAVDRL LMRSRLGGGG GAFGLGFGFG 180 FGGAHHLQAP PRCQAEGCKA DLSGAKHYHR RHKVCEYHAK ASVVATGGKQ QRFCQQCSRF 240 HVLSEFDEVK RSCRKRLAEH NRRRRKPPTT ASSKDAASPP ANKKPNCGSI TSSYSTDNKN 300 LNTAKSTMSS NTGSVISCLD QAAGNKQLQA RPTLTLGASP DRDQLHHQLR AMLQAQAAAG 360 GGHHHHQEQH FITSLQVHNN DGCGGGNSSS SNNNIILSCS SVCSSALPSA NGEVSDQNNG 420 NAGGMHNLFE VDFM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-28 | 193 | 266 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 249 | 266 | KRSCRKRLAEHNRRRRKP |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Zm.237 | 0.0 | meristem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G101499 | |||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in stems, leaf sheaths, and young panicles. {ECO:0000269|PubMed:16861571}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in stems, leaf sheaths, and young panicles. {ECO:0000269|PubMed:16861571}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00090 | SELEX | Transfer from AT1G02065 | Download |
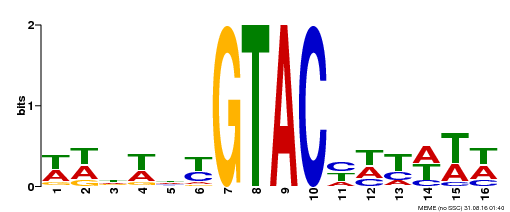
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G101499_P01 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AJ011616 | 0.0 | AJ011616.1 Zea mays mRNA for SBP-domain protein 3. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001105116.2 | 0.0 | SBP-domain protein 3 | ||||
| Swissprot | A2YFT9 | 3e-96 | SPL10_ORYSI; Squamosa promoter-binding-like protein 10 | ||||
| Swissprot | Q0DAE8 | 3e-96 | SPL10_ORYSJ; Squamosa promoter-binding-like protein 10 | ||||
| TrEMBL | A0A1D6LQH3 | 0.0 | A0A1D6LQH3_MAIZE; SBP-domain protein3 | ||||
| STRING | GRMZM2G101499_P01 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2375 | 35 | 93 | Representative plant | OGRP97 | 17 | 230 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G02065.1 | 9e-47 | squamosa promoter binding protein-like 8 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G101499_P01 |



