 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G087804_P01 | ||||||||
| Common Name | LOC542493 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 459aa MW: 49027.5 Da PI: 7.0936 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 86.8 | 2.1e-27 | 182 | 236 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k +++WtpeLH+rFv+aveqL G++kA+P++ile+m+ ++Lt+++++SHLQkYR++
GRMZM2G087804_P01 182 KRKVDWTPELHRRFVQAVEQL-GIDKAVPSRILEIMGTDCLTRHNIASHLQKYRSH 236
5799*****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.772 | 179 | 238 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.51E-17 | 182 | 239 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.9E-26 | 182 | 239 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 6.5E-27 | 182 | 236 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 5.4E-7 | 185 | 234 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 459 aa Download sequence Send to blast |
MLEVSTLRGP TSSGSKAEQH CGGGGGFVGD HHVVFPTSGD CFAMVDDNLL DYIDFSCDVP 60 FFDADGDILP DLEVDTTELL AEFSSTPPAD DLLAVAVFGA DDQPAAAVAQ EKPSSSLEQT 120 CGDDKGVAVA AARRKLQTTT TTTTTEEEDS SPAGSGANKS SASAEGHSSK KKSAGKNSNG 180 GKRKVDWTPE LHRRFVQAVE QLGIDKAVPS RILEIMGTDC LTRHNIASHL QKYRSHRKHL 240 MAREAEAATW AQKRHMYAPP APRTTTTTDA ARPPWVVPTT IGFPPPRFCR PLHVWGHPPP 300 HAAAAEAAAA TPMLPVWPRH LAPPRHLAPW AHPTPVDPAF WHQQYSAARK WGPQAAAVTQ 360 GTPCVPLPRF PVPHPIYSRP AMVPPPPSTT KLAQLHLELQ AHPSKESIDA AIGDVLVKPW 420 LPLPLGLKPP SLDSVMSELH KQGVPKIPPA AATTTGATG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 2e-14 | 182 | 234 | 5 | 57 | ARR10-B |
| 5lxu_A | 2e-14 | 187 | 238 | 6 | 57 | Transcription factor LUX |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G087804 | |||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in leaves. {ECO:0000269|PubMed:11340194}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that promotes chloroplast development. Acts as an activator of nuclear photosynthetic genes involved in chlorophyll biosynthesis, light harvesting, and electron transport (By similarity). {ECO:0000250}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
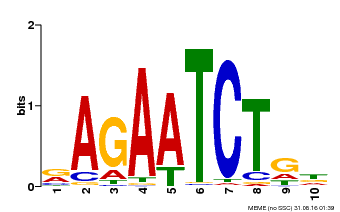
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G087804_P01 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. {ECO:0000269|PubMed:11340194}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF318579 | 0.0 | AF318579.1 Zea mays putative transcription factor GOLDEN 2 mRNA, complete cds. | |||
| GenBank | BT062838 | 0.0 | BT062838.1 Zea mays full-length cDNA clone ZM_BFb0374G22 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008673438.1 | 0.0 | golden plant 2 isoform X1 | ||||
| Swissprot | Q5NAN5 | 1e-110 | GLK2_ORYSJ; Probable transcription factor GLK2 | ||||
| TrEMBL | A0A1D6MES1 | 0.0 | A0A1D6MES1_MAIZE; Golden plant2 | ||||
| STRING | GRMZM2G087804_P03 | 0.0 | (Zea mays) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 7e-59 | GBF's pro-rich region-interacting factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G087804_P01 |
| Entrez Gene | 542493 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



