 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GRMZM2G025642_P01 | ||||||||
| Common Name | LOC100383060 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Tripsacinae; Zea
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 362aa MW: 39962.5 Da PI: 5.6877 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 175.2 | 1.8e-54 | 8 | 137 | 1 | 129 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk...kvkaeekewyfFskrdkkyatgkrknratksgyWkatgkd 89
+ppGfrFhPtdeelv +yL+kkv+++k++l +vi+++d+y++ePwdL+ ++e++ewyfFs +d+ky+tg+r+nrat +g+Wkatg+d
GRMZM2G025642_P01 8 VPPGFRFHPTDEELVGYYLRKKVAAQKIDL-HVIRDIDLYRIEPWDLQGhcgIGYEEQNEWYFFSYKDRKYPTGTRTNRATMAGFWKATGRD 98
69****************************.99**************943543334677********************************* PP
NAM 90 kevlskkgelvglkktLvfykgrapkgektdWvmheyrle 129
k+v++ k++ +g++ktLvfykgrap+g+ktdW+mheyrle
GRMZM2G025642_P01 99 KAVHD-KSRIIGMRKTLVFYKGRAPNGHKTDWIMHEYRLE 137
*****.999*****************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 8.11E-60 | 5 | 157 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 59.045 | 8 | 157 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.7E-28 | 9 | 136 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0048759 | Biological Process | xylem vessel member cell differentiation | ||||
| GO:1901348 | Biological Process | positive regulation of secondary cell wall biogenesis | ||||
| GO:1990110 | Biological Process | callus formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 362 aa Download sequence Send to blast |
MNSMESCVPP GFRFHPTDEE LVGYYLRKKV AAQKIDLHVI RDIDLYRIEP WDLQGHCGIG 60 YEEQNEWYFF SYKDRKYPTG TRTNRATMAG FWKATGRDKA VHDKSRIIGM RKTLVFYKGR 120 APNGHKTDWI MHEYRLETDE NAPPQEEGWV VCRAFKKRTA YPARMAWGDP SYAYREISAM 180 GAAAAAFVDP NAASYAQIRR QQSNKSARFK QEAELDGGAA AALLQYSSSH LVELPQLESP 240 SAPANKSQAA SASDEVVDAA DSGRRPGKKA RADEVATDWR ALDKFVASQL SPAAECGLEA 300 AAASSTAAAS NVGSQLDHVE DDDMAALLFL NSDGRDEAER WTGLLGPAAG DDGDFGLCVF 360 DK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 3e-49 | 5 | 161 | 12 | 172 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | GRMZM2G025642 | |||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Up-regulated during xylem vessel element formation. Expressed preferentially in procambial cells adjacent to root meristem. {ECO:0000269|PubMed:16103214, ECO:0000269|PubMed:25148240}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in root metaxylem pole and in shoot pre-procambium and procambium (PubMed:18445131). Present in root developing xylems (PubMed:16103214). Specifically expressed in vessels but not in interfascicular fibers in stems (PubMed:25148240). {ECO:0000269|PubMed:16103214, ECO:0000269|PubMed:18445131, ECO:0000269|PubMed:25148240}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the secondary wall NAC binding element (SNBE), 5'-(T/A)NN(C/T)(T/C/G)TNNNNNNNA(A/C)GN(A/C/T)(A/T)-3', in the promoter of target genes (By similarity). Involved in xylem formation by promoting the expression of secondary wall-associated transcription factors and of genes involved in secondary wall biosynthesis and programmed cell death, genes driven by the secondary wall NAC binding element (SNBE). Triggers thickening of secondary walls (PubMed:25148240). {ECO:0000250|UniProtKB:Q9LVA1, ECO:0000269|PubMed:25148240}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00266 | DAP | Transfer from AT2G18060 | Download |
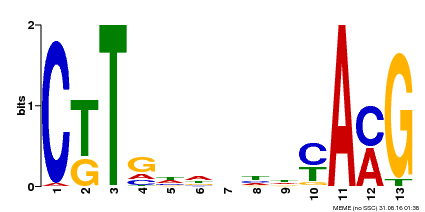
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | GRMZM2G025642_P01 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by chitin (e.g. chitooctaose) (PubMed:17722694). Accumulates in plants exposed to callus induction medium (CIM) (PubMed:17581762). {ECO:0000269|PubMed:17581762, ECO:0000269|PubMed:17722694}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT066067 | 0.0 | BT066067.1 Zea mays full-length cDNA clone ZM_BFb0225F19 mRNA, complete cds. | |||
| GenBank | BT083793 | 0.0 | BT083793.1 Zea mays full-length cDNA clone ZM_BFb0040P02 mRNA, complete cds. | |||
| GenBank | JN634081 | 0.0 | JN634081.1 Zea mays secondary wall NAC transcription factor 5 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001169207.1 | 0.0 | uncharacterized protein LOC100383060 | ||||
| Refseq | XP_008644372.1 | 0.0 | secondary wall NAC transcription factor 5 isoform X1 | ||||
| Swissprot | Q9SL41 | 1e-128 | NAC37_ARATH; NAC domain-containing protein 37 | ||||
| TrEMBL | C0PCP8 | 0.0 | C0PCP8_MAIZE; Secondary wall NAC transcription factor 5 | ||||
| TrEMBL | K0D9P4 | 0.0 | K0D9P4_MAIZE; NAC33 NAC type transcription factor (Fragment) | ||||
| STRING | GRMZM2G025642_P02 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3625 | 37 | 68 | Representative plant | OGRP17 | 15 | 800 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G18060.1 | 1e-115 | vascular related NAC-domain protein 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GRMZM2G025642_P01 |
| Entrez Gene | 100383060 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



