 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015892916.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 434aa MW: 46121.8 Da PI: 10.4614 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 42.9 | 1.1e-13 | 356 | 406 | 5 | 55 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkke 55
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L++ Nk+L+ + e ++
XP_015892916.1 356 RRQRRMIKNRESAARSRARKQAYTLELEAEVAKLKETNKELQRKQAECMEM 406
79************************************8776554444443 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.3E-11 | 352 | 418 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.611 | 354 | 399 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 7.5E-10 | 356 | 403 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 3.6E-13 | 356 | 405 | No hit | No description |
| CDD | cd14707 | 6.20E-23 | 356 | 410 | No hit | No description |
| Pfam | PF00170 | 4.6E-11 | 356 | 407 | IPR004827 | Basic-leucine zipper domain |
| PROSITE pattern | PS00036 | 0 | 359 | 374 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900057 | Biological Process | positive regulation of leaf senescence | ||||
| GO:1903648 | Biological Process | positive regulation of chlorophyll catabolic process | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 434 aa Download sequence Send to blast |
MGSHMNFKNF GDVPLMDGGG GGGGKATGNF TLARQPSIYS LTFDELQNTL GGIGKDFGSM 60 NMDELLKNIW TAEETQIVTS TAGAGGGTNP GGNLQRQGSL TLPRTLSQKK VDEVWRDLLK 120 EGGGGTDTIG GGGGGGGGGG CGGATNLPRQ QTLGEMTLEE FLVRAGVVRE EAQQIGRQAN 180 SGFYNEFYRS NNNNGLALGY QQPGRNNVLL DNQVAQKNIN SAPNQPPSLA RSGGGIRSSQ 240 QHMQQPPSQQ QPLLPKPANI AFAPTTHLVS NGQATRGIVS RIAEPAMNTS LVQAGGIGMA 300 GLGAGGVTVA AGSPGSHISS DVISKSSVDT PSFSPVPYVF SRGRKSGGAL EKVVERRQRR 360 MIKNRESAAR SRARKQAYTL ELEAEVAKLK ETNKELQRKQ AECMEMQKNQ ILEKMNRQNG 420 NKRQCLRRTL TGPW |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Functions as transcriptional activator in the ABA-inducible expression of rd29B. Binds specifically to the ABA-responsive element (ABRE) of the rd29B gene promoter. {ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16463099}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
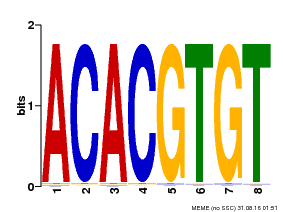
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015892916.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA) and cold. {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015892916.1 | 0.0 | bZIP transcription factor 46-like | ||||
| Swissprot | Q9M7Q2 | 1e-113 | AI5L7_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| TrEMBL | A0A2I4EER7 | 0.0 | A0A2I4EER7_JUGRE; ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| STRING | XP_004294799.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1962 | 34 | 81 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G19290.1 | 2e-89 | ABRE binding factor 4 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



