 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015882347.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1454aa MW: 162472 Da PI: 8.8928 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.6 | 0.0002 | 1363 | 1386 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirtH 23
+Cp Cgk F ++ +L++H r+H
XP_015882347.1 1363 VCPvkGCGKKFFSHKYLVQHRRVH 1386
69999*****************99 PP
| |||||||
| 2 | zf-C2H2 | 11.9 | 0.00071 | 1422 | 1448 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
XP_015882347.1 1422 YVCAepGCGQTFRFVSDFSRHKRKtgH 1448
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 9.8E-17 | 20 | 61 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.603 | 21 | 62 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 2.6E-14 | 22 | 55 | IPR003349 | JmjN domain |
| PROSITE profile | PS51184 | 33.982 | 192 | 361 | IPR003347 | JmjC domain |
| SMART | SM00558 | 2.0E-49 | 192 | 361 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 2.17E-27 | 206 | 375 | No hit | No description |
| Pfam | PF02373 | 1.2E-36 | 225 | 344 | IPR003347 | JmjC domain |
| SMART | SM00355 | 28 | 1339 | 1361 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.0037 | 1362 | 1386 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 5.88E-6 | 1362 | 1398 | No hit | No description |
| PROSITE profile | PS50157 | 12.549 | 1362 | 1391 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 8.4E-6 | 1363 | 1385 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1364 | 1386 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 5.3E-9 | 1386 | 1414 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0017 | 1392 | 1416 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.637 | 1392 | 1421 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1394 | 1416 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.57E-9 | 1402 | 1446 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.3E-9 | 1415 | 1445 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.62 | 1422 | 1448 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.24 | 1422 | 1453 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1424 | 1448 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1454 aa Download sequence Send to blast |
MAASAEPPQE VFSWLKTLPS APEYHPTLAE FQDPISYIFK IEKEASKYGI CKIVPPVPPS 60 PRKTAMANLH KSLAARSAFS DSSNPKSQPT FTTRQQQIGF CPRKPRPVQR PVWQSGEYYT 120 FSQFETRAKN FEKTFLKKCT KKGGLSPLEI ETLYWKATVD KPFSVEYAND MPGSAFVPVS 180 TKKSREAGEG VTLGETAWNM RAVSRAKGSL LRFMKEEIPG VTSPMVYIAM MFSWFAWHVE 240 DHDLHSLNYL HMGAGKTWYG VPREAAVAFE EVVRVHGYGG EVNPLVTFAI LGEKTTVMSP 300 EVFISAGIPC CRLVQNPGEF VVTFPRAYHT GFSHGFNCGE AANIATPEWL GVAKDAAIRR 360 ASINYPPMVS HYQLLYDLAL ALCSRVPASI TAEPRSSRLK DKKKGEGETV VKEQFVQDVI 420 HNNDLLHVLG KGSPIVLLPR SSLDISVCSK LRVGSQLRVN TTFPVGLCNS GGEMKSSGSL 480 ISDDLMMMDT KQEIKQVKGF YLVKGKFASL CEGSRIPSLS GNDSQFSLNA KSVGIEGENN 540 SGNDGLSDQR LFSCVTCGIL SFSCVAIIQP REPAARYLMS ADCSFFNDWV VGSGVASDVF 600 TVANADPIAQ NTCTGLVEKN GPDSLYDVPV QSVNEDKSNE VVSNAEKHEP ASALGLLALN 660 YGNSSDSEDD QVDPNVSVYG AETILPNRAS GIYQYDHSSS TLQDRHVDAS GLQGQSSCRL 720 YSGGGLASLN VDLNMDNCTV DVDSDNLAST KSNGLVGQFR DPMNVLHACS SGSYDAETTK 780 FGKGITPVKN ANMPFAPRCD EESSRMHVFC LDHAVEVEQQ LRQIGGVDIL LLCHPDYPKI 840 EAEAKVIAED LGIGCLWNAH TFRKATKEDE ERIQAALDSE EAIPKNGDWA VKLGINLFYS 900 ASLSRSPLYS KQMPYNSVIY NAFGRSSPAV SPTRSDGYVR RSGKQKKVVA GKWCGKVWMS 960 SQVHPFLAKR DPEEEEDEER GFHTWPIPDE RLERKPDTSR RNDSTMITRK YVRKRKMAVE 1020 SGLNKKAKCI QKEDAVSGNS MNDDSHQQQR RLPRSKEVCI KRGSTKKIKC IETEEAVSDD 1080 SIDDNSDEQH KRTHRRNQAN YIERDDIVSD DSLGVDSHHQ NKQIFRGKQA KHTGREDVVS 1140 DDSLDSNSCL LNRRIPRSKQ SKFFGREDSV SDDFYGDNAQ QQQRRIQSKQ AKSMEREDEI 1200 LDEPVEDNSH QQHKRILRNK RTKSAIQQQK MKRETSRNVK QAISKPVKQG TRRQEKMKQQ 1260 TPRLRNNPSE QNMFYTNAEE ELEGGPSTRL RKRIPKPLKV AGAKRKEQQQ PKSKKKVKKA 1320 STGKAQGGHN DARDEEAEYL CDLEGCTMSF GTKQELVLHK RNVCPVKGCG KKFFSHKYLV 1380 QHRRVHMDDR PLRCPWKGCK MTFKWAWART EHIRVHTGAR PYVCAEPGCG QTFRFVSDFS 1440 RHKRKTGHSA KKGR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 6e-78 | 14 | 382 | 8 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 6e-78 | 14 | 382 | 8 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1009 | 1015 | KYVRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
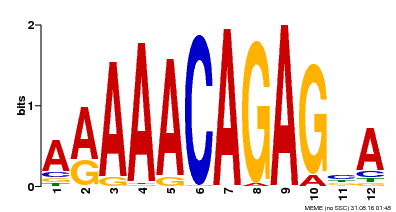
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015882347.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015882347.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A2P5BA48 | 0.0 | A0A2P5BA48_PARAD; TFIIH C1-like domain containing protein | ||||
| STRING | XP_010101942.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5648 | 32 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



