 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_015874519.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rhamnaceae; Paliureae; Ziziphus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 540aa MW: 60400 Da PI: 6.2904 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 39.2 | 1.6e-12 | 490 | 536 | 3 | 46 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHT...TTS-HHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmg...kgRtlkqcksrwqk 46
+W+ E+e l d+vk++G+g+Wk+I ++ + Rt+ ++k++w++
XP_015874519.1 490 KWSLVEEETLRDGVKKFGKGNWKLILNCYRdvfEERTEVDLKDKWRN 536
8********************************9************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.162 | 483 | 540 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.7E-8 | 487 | 540 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.95E-14 | 490 | 539 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-11 | 490 | 538 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.2E-9 | 490 | 536 | IPR001005 | SANT/Myb domain |
| CDD | cd11660 | 4.12E-18 | 491 | 539 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009901 | Biological Process | anther dehiscence | ||||
| GO:0010152 | Biological Process | pollen maturation | ||||
| GO:0043067 | Biological Process | regulation of programmed cell death | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 540 aa Download sequence Send to blast |
MDVDVSRWIL EFLLRKCDDD GVVKKALKVI PFPNDESRLK KVVLLRAIES EVSEASVSEK 60 ILENLEMLEE LDQREGTDTL DSMKAAYCAV ALECTAKYLS ASKPGKYFEA VRRIWRGRVS 120 HMERSGKSRL LTEELKKNGE EVEAAIWDAN VSKKLCRMNT RNEALGLLSV YLGEAWALMG 180 PSFIEWAARL SIKDKGVDGS GVEEGIGDGV DRDRGDVDNG GNAKDGGHEV GAQSNVGVEV 240 PEQVVVDSPE RCAESGLNGL LELAVQFENR VDGNGNVSDH NPAAKKEVTT PKDKETRKEN 300 VGLRCKHVAA HRRHRGPARI TNGGDLDPEA LCNKDDSLSS ADVNKARQSL KSSRMELLAV 360 VTDPLPEAIQ RSEIIISNLA TKNVNHEPSL QNEGRKDLDE SNSSVDKNAE TIQPSDINVG 420 DPSCSHQKNN VPRPTLMERN STAQTYEWDD SIDGSPERNR LHLPSPKRKI SSPLQKYEMT 480 KIAKRRKIKK WSLVEEETLR DGVKKFGKGN WKLILNCYRD VFEERTEVDL KDKWRNMSRY |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 466 | 486 | KRKISSPLQKYEMTKIAKRRK |
| 2 | 466 | 489 | KRKISSPLQKYEMTKIAKRRKIKK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00052 | PBM | Transfer from AT1G15720 | Download |
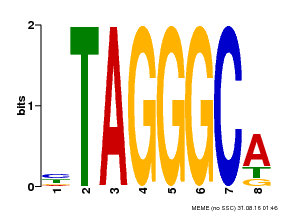
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_015874519.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015874519.1 | 0.0 | uncharacterized protein LOC107411450 isoform X1 | ||||
| TrEMBL | A0A2P5AM89 | 1e-176 | A0A2P5AM89_PARAD; GAMYB transcription factor | ||||
| TrEMBL | A0A2P5CYN7 | 1e-176 | A0A2P5CYN7_TREOI; GAMYB transcription factor | ||||
| STRING | XP_010108474.1 | 1e-155 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2824 | 32 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G06910.1 | 3e-15 | TRF-like 7 | ||||



