 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00071.1.g02550.1.sm.mkhc | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 612aa MW: 68362.7 Da PI: 4.8335 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 168.9 | 1.7e-52 | 9 | 142 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.k.kvkaeekewyfFskrdkkyatgkrknra 77
lp GfrFhPtdeelv +yLk k++g+ +++ evi+e+d++k+ePwdLp k +++e+ ew+fF+++d+ky++g+r+nra
Zjn_sc00071.1.g02550.1.sm.mkhc 9 LPLGFRFHPTDEELVRHYLKGKITGQINSEVEVIPEIDVCKCEPWDLPdKsLIRSEDPEWFFFAPKDRKYPNGSRSNRA 87
699************************9999****************96446677888********************* PP
NAM 78 tksgyWkatgkdkevlsk....kgelvglkktLvfykgrapkgektdWvmheyrl 128
t++gyWkatgkd+ + sk k++++g+kktLvf+kgrapkge+t W+mheyr
Zjn_sc00071.1.g02550.1.sm.mkhc 88 TEAGYWKATGKDRVIRSKgdkrKQHVIGMKKTLVFHKGRAPKGERTGWIMHEYRT 142
*****************988767777***************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 2.48E-58 | 3 | 163 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 56.524 | 9 | 163 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.5E-27 | 10 | 141 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0071470 | Biological Process | cellular response to osmotic stress | ||||
| GO:1900426 | Biological Process | positive regulation of defense response to bacterium | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005789 | Cellular Component | endoplasmic reticulum membrane | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 612 aa Download sequence Send to blast |
MTVMELKALP LGFRFHPTDE ELVRHYLKGK ITGQINSEVE VIPEIDVCKC EPWDLPDKSL 60 IRSEDPEWFF FAPKDRKYPN GSRSNRATEA GYWKATGKDR VIRSKGDKRK QHVIGMKKTL 120 VFHKGRAPKG ERTGWIMHEY RTTEPEFECG EQGSYVLYRL FCKQEEKNER LTPEEMDRSG 180 YSPTTPRSSP NNIEANEEAF TPLNKESPES TLHDSPTNLP NSVETHALSL TRWLVDRTDN 240 SAANAVNVSK TPLNGHIDGI AKVDPSVGGV TQLIDSQQQH NGDPNHFAAG SAPILQHEEN 300 GFFNFQQGMM GFDGNVNAPD AIDEFLNQAL TDPDEHSSTT SKVQYDSDPG IMAPEFGNHG 360 IMQGELPDDQ AWWAGLNFDL DEPNSLLPYD TTDPDVLSMD SGAESLHDLF NGWSTEPAVQ 420 GTGITIMPRQ SQSNVQQNNL FLRQGIANRR LRLQEYLSPN VESATNVDCE DKASCTVTSD 480 YLDEATEEST AEKGEPYYED DAESTGNTIR SRHPAPSSSA TSSFTQQGTA VRRLRLQSDL 540 KTDALKAVSE SKSVPRRRRT SEKNDEITKQ EDGFDSLVRA PGKKRGFQSH LFWLVLSVAV 600 LLLLCVGVYG WA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 3e-45 | 8 | 164 | 16 | 166 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 3e-45 | 8 | 164 | 16 | 166 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 3e-45 | 8 | 164 | 16 | 166 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 3e-45 | 8 | 164 | 16 | 166 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 3e-45 | 8 | 164 | 19 | 169 | NAC domain-containing protein 19 |
| 3swm_B | 3e-45 | 8 | 164 | 19 | 169 | NAC domain-containing protein 19 |
| 3swm_C | 3e-45 | 8 | 164 | 19 | 169 | NAC domain-containing protein 19 |
| 3swm_D | 3e-45 | 8 | 164 | 19 | 169 | NAC domain-containing protein 19 |
| 3swp_A | 3e-45 | 8 | 164 | 19 | 169 | NAC domain-containing protein 19 |
| 3swp_B | 3e-45 | 8 | 164 | 19 | 169 | NAC domain-containing protein 19 |
| 3swp_C | 3e-45 | 8 | 164 | 19 | 169 | NAC domain-containing protein 19 |
| 3swp_D | 3e-45 | 8 | 164 | 19 | 169 | NAC domain-containing protein 19 |
| 4dul_A | 3e-45 | 8 | 164 | 16 | 166 | NAC domain-containing protein 19 |
| 4dul_B | 3e-45 | 8 | 164 | 16 | 166 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00108 | PBM | Transfer from AT4G35580 | Download |
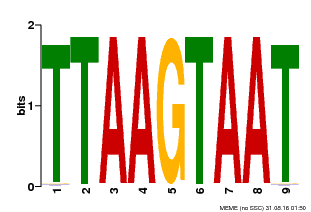
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002445085.1 | 0.0 | protein NTM1-like 9 isoform X2 | ||||
| Refseq | XP_025820792.1 | 0.0 | protein NTM1-like 9 | ||||
| TrEMBL | A0A0A9CTY0 | 0.0 | A0A0A9CTY0_ARUDO; Uncharacterized protein | ||||
| STRING | Sb07g003900.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5172 | 38 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G35580.1 | 6e-71 | NAC transcription factor-like 9 | ||||



