 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00043.1.g04860.1.am.mkhc | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 686aa MW: 74300 Da PI: 8.7951 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.1 | 1.3e-12 | 536 | 582 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI +Lq
Zjn_sc00043.1.g04860.1.am.mkhc 536 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAITYITDLQ 582
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 4.9E-49 | 131 | 316 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.831 | 532 | 581 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.31E-18 | 535 | 597 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 5.82E-15 | 535 | 586 | No hit | No description |
| Pfam | PF00010 | 4.3E-10 | 536 | 582 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 9.6E-18 | 536 | 594 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.9E-16 | 538 | 587 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 686 aa Download sequence Send to blast |
RAPEIHSRIR EPPAPSALIP CIRRHPRTHG VLSPCPGARL HTAGRDLATR LALDSAAPAS 60 IRAQGPQSGI MVMKMEIEDD AAIGGGTGGI WTEEDRALGA AVLGADAFSY LTKGGGAISQ 120 GLVSASLSVD MQNKLQELVE SDRPGAGWNY AIYWQLSRTK SGDFVLGWGD GSCREPCDGE 180 VGAAASAGND DTKQRMRKRV LQRLHIAFGG VDEEDYALGI DQVTDTEMFF LASMYFAFPR 240 RTGGPGQVFA AGIPLWIPNS ERKVYPTNYC YRGFLADAAG FRTIVLVPFE GGVLELGSMQ 300 HIAESSDAIQ NIRSVFSGTS NNKAAAQKPE GNGPAATERS PSLAKIFGKD LNLGRPSAGP 360 VAGVSKVDER SWEQRSAAGG GSLLPNVQKG LQNFTWSQAR GLNSHQQKFG NGVLIVSNEA 420 AQRSNGATDN SSATQFQLQK VLPPQKLQLQ KLPQIQKTPP MVNQQQLQPQ APRQIDFSTG 480 SSSKAGVLVT RTSGLDGESA EVDGLCKEEG PAPAVEDRRP RKRGRKPANG REEPLNHVEA 540 ERQRREKLNQ RFYALRAVVP NISKMDKASL LGDAITYITD LQKKLKEMET ERERFLESGL 600 VGGMPRPEVD IQVVQDEVLV RVMSPMENHP VKRVFQAFED AEVRIGESKV TGNNGMVVHS 660 FTIKCPGSEQ QTREKVIAAM SRAMNS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 7e-28 | 530 | 592 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 7e-28 | 530 | 592 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 7e-28 | 530 | 592 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 7e-28 | 530 | 592 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 7e-28 | 530 | 592 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 7e-28 | 530 | 592 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 7e-28 | 530 | 592 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 7e-28 | 530 | 592 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 517 | 525 | RRPRKRGRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
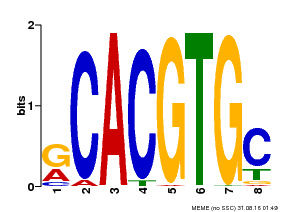
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025816895.1 | 0.0 | transcription factor bHLH13-like | ||||
| TrEMBL | A0A3L6SNM9 | 0.0 | A0A3L6SNM9_PANMI; Transcription factor bHLH13-like | ||||
| STRING | Pavir.Ea00030.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5174 | 38 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 1e-68 | bHLH family protein | ||||



