 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00027.1.g03830.1.sm.mkhc | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 465aa MW: 51933.7 Da PI: 4.8958 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 112.1 | 3.9e-35 | 21 | 112 | 2 | 102 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEE CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiwe 80
Fl k+ye++ed+++++++sw g sfvv+++ +f++++LpkyFkh+nf+SF+RQLn+YgF+k++ e+ we
Zjn_sc00027.1.g03830.1.sm.mkhc 21 FLIKTYEMVEDPATNHVVSWGPGGASFVVWNPPDFSQELLPKYFKHNNFSSFIRQLNTYGFRKIDPER---------WE 90
9****************************************************************999.........** PP
EEESXXXXXXXXXXXXXXXXXX CS
HSF_DNA-bind 81 FkhksFkkgkkellekikrkks 102
F+++ F +g+++ll++i+r+k
Zjn_sc00027.1.g03830.1.sm.mkhc 91 FANEDFIRGHTHLLKNIHRRKP 112
*******************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46785 | 2.58E-36 | 16 | 110 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Gene3D | G3DSA:1.10.10.10 | 3.4E-39 | 17 | 110 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 2.0E-62 | 17 | 110 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 8.7E-20 | 21 | 44 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 3.8E-31 | 21 | 110 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 8.7E-20 | 59 | 71 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 60 | 84 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 8.7E-20 | 72 | 84 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 465 aa Download sequence Send to blast |
MEGSQSRSGG GGGGSSSPPP FLIKTYEMVE DPATNHVVSW GPGGASFVVW NPPDFSQELL 60 PKYFKHNNFS SFIRQLNTYG FRKIDPERWE FANEDFIRGH THLLKNIHRR KPVHSHSVQN 120 QVNTLAESER REYEDEIKRL KYEKSLLLAD LQKQDQQRCL TNWQMQSLED RLMQMEQRQK 180 NIVASLCDIL QRNGAVASPL SLQTDHFNFS KKRRVPKIGF FADDDAAVEE ERQASFLQMM 240 GAVPKSPSVF PVQQHPVNAE PFDRMETSLV SLEKFIQRAS DADASPEDMF AIADAPSLAV 300 ALGEVQSAPM ETGINLQSST GQVHSSAGFA ESSCYASSPM LPLADLREDA PRTAKVDINS 360 DTTSTDTSQD EATTETGGSL SQEPAKVNDV FWERFLTETP KFGRAGEADS GRQEDECKSE 420 PAEVKEDEKV AVDCSSLHRY KVVDQITEQM GQLASAEYAV PHHLR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 1e-25 | 9 | 110 | 17 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 1e-25 | 9 | 110 | 17 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 1e-25 | 9 | 110 | 17 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds DNA of heat shock promoter elements (HSE). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
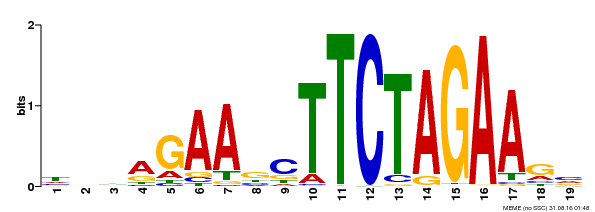
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat stress. {ECO:0000269|PubMed:12032317}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025810280.1 | 0.0 | heat stress transcription factor A-4d-like | ||||
| Refseq | XP_025810281.1 | 0.0 | heat stress transcription factor A-4d-like | ||||
| Swissprot | Q93VB5 | 1e-168 | HFA4D_ORYSJ; Heat stress transcription factor A-4d | ||||
| TrEMBL | A0A2S3HA79 | 0.0 | A0A2S3HA79_9POAL; Uncharacterized protein | ||||
| STRING | Si022026m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3296 | 37 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18880.1 | 3e-66 | heat shock transcription factor A4A | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



