 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00018.1.g07120.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 703aa MW: 75534.2 Da PI: 9.1788 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.2 | 3.2e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd+ lv +v+++G+g+W++++ g+ R+ k+c++rw +yl
Zjn_sc00018.1.g07120.1.am.mk 14 KGPWTPEEDLMLVSYVQEHGPGNWRAVPTNTGLMRCSKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 47.3 | 4.7e-15 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T +E++l++++ +lG++ W++Ia++++ Rt++++k++w+++l
Zjn_sc00018.1.g07120.1.am.mk 67 RGNFTDQEEKLIIHLQVLLGNR-WAAIASYLP-ERTDNDIKNYWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 9.5E-24 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 24.935 | 9 | 65 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.54E-29 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 7.5E-14 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.8E-16 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.76E-11 | 16 | 61 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-23 | 65 | 117 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.831 | 66 | 116 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.3E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 7.3E-14 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 6.58E-11 | 69 | 112 | No hit | No description |
| ProDom | PD125774 | 5.0E-24 | 614 | 693 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| Pfam | PF04770 | 2.3E-27 | 634 | 686 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| TIGRFAMs | TIGR01566 | 2.4E-20 | 635 | 684 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| PROSITE profile | PS51523 | 24.361 | 636 | 685 | IPR006456 | ZF-HD homeobox protein, Cys/His-rich dimerisation domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0080167 | Biological Process | response to karrikin | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 703 aa Download sequence Send to blast |
MGRPPCCDKV GVKKGPWTPE EDLMLVSYVQ EHGPGNWRAV PTNTGLMRCS KSCRLRWTNY 60 LRPGIKRGNF TDQEEKLIIH LQVLLGNRWA AIASYLPERT DNDIKNYWNT HLKKKLKKMQ 120 AGESGGGDQD GVRVSESAGA GDRVGGEKRP TVHKGQWERR LQTDIHTARQ ALREALSLEP 180 SPPAKMAPLP TPAGSATYAS SAENIARLLE GWLRPGSGCG KGPETSGSTF TTGTTQQRPQ 240 CSGEGAASAS ESHSGGTAAM AVQTPECSTE TSKMVGAAGA GAPPAFSMLE NWLLDDGMAH 300 GEVGLIDVAW LGALVRDSGT AMATHACYSY SSMDMHESSK ASHRKKLEKK KARVKSNATR 360 DPGGRDGRSA TPCGLGNAQG PDSMPAPTQA GRPQGATALL RAPCIMQACR GTPNLIQSSC 420 REEGRKEGRT HAVEGRWRVP SAHLVRGLNA PALPAPSLID LKALSPLITA APLRSRNGME 480 LNGPPEADPG DGIRRRRDGE TSAPPTDKRA RTYCCRPAGS VRSIDRNSRG AVGVRSGTVR 540 SILSDRMEVI GARYERPAVA SPTRRISPVS HDDVGGGGGD SYLPVVHSKQ GLHSPCAETR 600 LWLAGWEEEG RLTMKRLLIL RRCEPIVRFT CCSVRYGECR RNHAATTGGY SVDGCREFIA 660 EGEEGTGGAL KCAACGCHRS FHRRVQVYEV SWDYAASDSS STE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1mse_C | 6e-25 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| 1msf_C | 6e-25 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00395 | DAP | Transfer from AT3G47600 | Download |
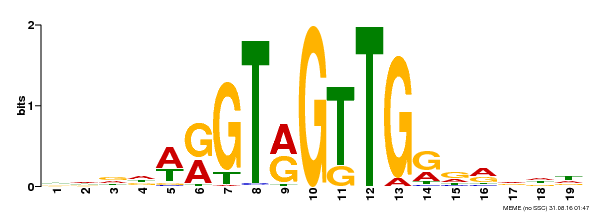
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JF951914 | 1e-157 | JF951914.1 Aegilops tauschii clone TaMYB31 R2R3-MYB protein mRNA, complete cds. | |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP229 | 38 | 296 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G47600.1 | 2e-94 | myb domain protein 94 | ||||



