 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zjn_sc00013.1.g05960.1.sm.mkhc | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 344aa MW: 38943.8 Da PI: 9.4882 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 163.6 | 7e-51 | 37 | 165 | 2 | 128 |
NAM 2 ppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.kkvkaeekewyfFskrdkkyatgkrknratk 79
pGfrFhPtdeelv++yL++kv++k+l++ e+ike+diyk++PwdLp +++ ++ekewyfF+ r +ky+++ r+nr+t
Zjn_sc00013.1.g05960.1.sm.mkhc 37 LPGFRFHPTDEELVTFYLRRKVARKPLSI-EIIKEMDIYKHDPWDLPkASTVGGEKEWYFFCLRGRKYRNSIRPNRVTG 114
69***************************.89***************445556889*********************** PP
NAM 80 sgyWkatgkdkevlsk..kgelvglkktLvfykgrapkgektdWvmheyrl 128
sg+Wkatg d++++s+ +g+ +glkk+Lv+y g+a kg+ktdW+mhe+rl
Zjn_sc00013.1.g05960.1.sm.mkhc 115 SGFWKATGIDRPIYSAtnSGKSIGLKKSLVYYCGSAGKGTKTDWMMHEFRL 165
**************99878889***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51005 | 55.318 | 36 | 193 | IPR003441 | NAC domain |
| SuperFamily | SSF101941 | 5.89E-56 | 36 | 193 | IPR003441 | NAC domain |
| Pfam | PF02365 | 3.7E-25 | 38 | 165 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005992 | Biological Process | trehalose biosynthetic process | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006561 | Biological Process | proline biosynthetic process | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0010120 | Biological Process | camalexin biosynthetic process | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 344 aa Download sequence Send to blast |
MVAPKEFSRE AVSMDKVKSD GEALMAAGDG EEDDMVLPGF RFHPTDEELV TFYLRRKVAR 60 KPLSIEIIKE MDIYKHDPWD LPKASTVGGE KEWYFFCLRG RKYRNSIRPN RVTGSGFWKA 120 TGIDRPIYSA TNSGKSIGLK KSLVYYCGSA GKGTKTDWMM HEFRLPPAAG ADVNASPSMQ 180 EAEVWTICRI FKRNITYKRR QQQQQQQQVW RQPAGGGGKA PRPADSSSNT GSFESDGRDD 240 YMNRLPGAAP GISHHQQLSH QVNMLNGGGR SGFFRDNQLH SHQWLNFNSL TMPELQEQKP 300 QVNPPAMTIA FQQNHATNEF YRDGYLEDIA RYPSELKKGF HPSS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 3e-49 | 29 | 199 | 8 | 174 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
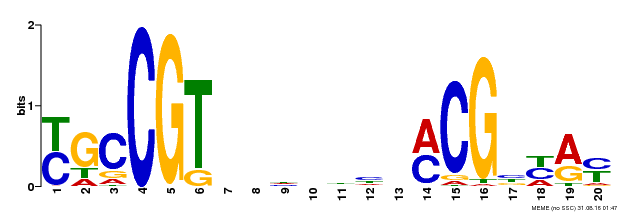
| UniProt | Transcription factor that binds to the 5'- RRYGCCGT-3' consensus core sequence. Central longevity regulator. Negative regulator of leaf senescence. Modulates cellular H(2)O(2) levels and enhances tolerance to various abiotic stresses through the regulation of DREB2A. {ECO:0000269|PubMed:22345491}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00310 | DAP | Transfer from AT2G43000 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by H(2)O(2), paraquat, ozone, 3-aminotriazole and salt stress. {ECO:0000269|PubMed:22345491}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK073539 | 1e-161 | AK073539.1 Oryza sativa Japonica Group cDNA clone:J033041C24, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025795725.1 | 0.0 | transcription factor JUNGBRUNNEN 1-like | ||||
| Swissprot | Q9SK55 | 2e-81 | NAC42_ARATH; Transcription factor JUNGBRUNNEN 1 | ||||
| TrEMBL | A0A2S3IHK9 | 1e-180 | A0A2S3IHK9_9POAL; Uncharacterized protein | ||||
| STRING | Si036447m | 1e-180 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1191 | 38 | 129 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43000.1 | 2e-83 | NAC domain containing protein 42 | ||||



