 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vradi07g18820.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1110aa MW: 125061 Da PI: 5.5601 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 161.5 | 1.5e-50 | 10 | 116 | 2 | 108 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
++ ++rwl++ ei+aiL n++k++++ e++t p+sgsl+L++rk++ryfrkDG++w+kkkdgktvrE+he+LK g+v+vl+cyYah+ee+++fq
Vradi07g18820.1 10 VEAQHRWLRPAEICAILSNYKKFRIAPEPATMPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVREAHERLKAGSVDVLHCYYAHGEEDESFQ 103
5679****************************************************************************************** PP
CG-1 96 rrcywlLeeelek 108
rr+ywlLee+l+
Vradi07g18820.1 104 RRTYWLLEERLRY 116
********99665 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 70.111 | 5 | 134 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.0E-61 | 8 | 160 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.0E-43 | 11 | 118 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF01833 | 1.9E-5 | 526 | 602 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 4.67E-15 | 526 | 611 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:2.60.40.10 | 2.0E-4 | 527 | 600 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF48403 | 2.49E-19 | 706 | 822 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 5.21E-16 | 708 | 820 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 7.3E-19 | 711 | 826 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 19.863 | 711 | 832 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 3.5E-9 | 733 | 828 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.965 | 761 | 793 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.002 | 761 | 790 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 390 | 800 | 829 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.38E-7 | 928 | 983 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.32 | 933 | 955 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.407 | 935 | 963 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0045 | 935 | 954 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.038 | 956 | 978 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.62 | 957 | 981 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 9.8E-4 | 959 | 976 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1110 aa Download sequence Send to blast |
MGIDIKQIIV EAQHRWLRPA EICAILSNYK KFRIAPEPAT MPPSGSLFLF DRKVLRYFRK 60 DGHNWRKKKD GKTVREAHER LKAGSVDVLH CYYAHGEEDE SFQRRTYWLL EERLRYFLGV 120 NFLKHGGTKA NYTCAKENEE SLPCAQQTDK IMLKAEMDTS FSSVLHPNSY QVPSQTMDSS 180 MNSAQVSEYE EAESAFNGHA SSEFYSFLEL QRPVEKIIDQ AADSYSPHPL IRKSITNMSF 240 SIETGTDEQK KLPVIAETNY VSLTQDRKFI DILSGGLTYE SLKPLGFSSW EDILENNGES 300 QHMPFQPLFP EMQPDNMAVN SNFCQGYDIM VPHLTTSIAK LHDNGSLIQA EGSWQGYSVD 360 SLRVSSWPID NVHSSSTCEV SCSNCEHEVN EVDIQKSLQQ SLLHQHKQDK VLMLNDPQEI 420 MLDYLKSDFE ASRTLDGIED PRFTFKRTQL DGSPAEEGLK KLDSFNQWMS KELGDVEESN 480 KPSTSGAYWD TVESESEVGS TTIPSQGHLD TYVLDPSVSN EQLFSIIDYS PSWAFEGSEV 540 KIIISGRFLR SQQEAEQCKW SCMFGEVEVP AVILAKDVLC CHTPQHKAGR VPFYVTCSNR 600 LACSEVREFD FQVNCTQEVN TAGEDRASIF HTLSRRFGEL LSLGHAVPQN SDSISGNEKS 660 QLRSKISSLL RGEDDVWDKL LELTQENEFS TEDLQEQLLQ NLLKDKLHAW LLQKITDDGK 720 GPNVLDEGGQ GVLHFAAALG YDWALEPTLV AGVNVNFRDI NGWTALHWAA FYGRERTVAF 780 LISLGAAPGL LTDPCPEHPS GRTPADLAST NGHKGIAGYL AESSLSAQLT TLDLNKDVGE 840 NSGNKVVQRI QNTGPVNDLD GLSYEQSLKD SLAAVCNATQ AAARIHQVFR MQSFQRKQLK 900 EFGDDKFGIS DERALSLVKK NGKSHKSGSR DEPVHAAAIR IQNKFRGWKG RKDFLMIRQR 960 IVKIQAHVRG HQVRKNCGKI IWTVGILEKV ILRWRRKGSG LRGFKAEANS EVTMIEDVTS 1020 SEDDYDVLKE GRKQTEQRLQ KALARVKSMI QYPEARDQYH RVLNVVTQIQ ENQVNHDSSC 1080 NNSEEIRDFN NLTDLEALLD EDIFMPTAT* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
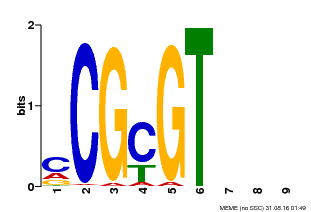
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vradi07g18820.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015034 | 0.0 | AP015034.1 Vigna angularis var. angularis DNA, chromosome 1, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014508804.1 | 0.0 | calmodulin-binding transcription activator 3 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A1S3USB7 | 0.0 | A0A1S3USB7_VIGRR; calmodulin-binding transcription activator 3 | ||||
| STRING | XP_007159660.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8284 | 27 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



