 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT2G22300.1 | ||||||||
| Common Name | CAMTA3, CMTA3, SR1, T26C19.4 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1032aa MW: 116111 Da PI: 5.2453 | ||||||||
| Description | signal responsive 1 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 182.5 | 4.8e-57 | 20 | 136 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcy 99
+e ++rwl++ ei++iL+n++++++++e++t+p+sgs+++++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+v+vl+cyYah+++n++fqrr+y
AT2G22300.1 20 SEaRHRWLRPPEICEILQNYQRFQISTEPPTTPSSGSVFMFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKAGSVDVLHCYYAHGQDNENFQRRSY 117
4449********************************************************************************************** PP
CG-1 100 wlLeeelekivlvhylevk 118
wlL+eel++iv+vhylevk
AT2G22300.1 118 WLLQEELSHIVFVHYLEVK 136
****************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.708 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.4E-85 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.0E-49 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 7.5E-5 | 463 | 551 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 5.76E-17 | 465 | 550 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 5.8E-4 | 465 | 550 | IPR002909 | IPT domain |
| SuperFamily | SSF48403 | 2.8E-15 | 636 | 756 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.02E-11 | 641 | 753 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 3.9E-15 | 641 | 756 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 16.149 | 644 | 753 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.164 | 694 | 726 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 3.72E-8 | 849 | 902 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.92 | 851 | 873 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.029 | 853 | 881 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0041 | 854 | 872 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.023 | 874 | 896 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.066 | 875 | 899 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 7.8E-4 | 877 | 896 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0045944 | Biological Process | positive regulation of transcription from RNA polymerase II promoter | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0001077 | Molecular Function | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000005 | anatomy | cultured plant cell | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1032 aa Download sequence Send to blast |
MAEARRFSPV HELDVGQILS EARHRWLRPP EICEILQNYQ RFQISTEPPT TPSSGSVFMF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHER LKAGSVDVLH CYYAHGQDNE NFQRRSYWLL 120 QEELSHIVFV HYLEVKGSRV STSFNRMQRT EDAARSPQET GDALTSEHDG YASCSFNQND 180 HSNHSQTTDS ASVNGFHSPE LEDAESAYNQ HGSSTAYSHQ ELQQPATGGN LTGFDPYYQI 240 SLTPRDSYQK ELRTIPVTDS SIMVDKSKTI NSPGVTNGLK NRKSIDSQTW EEILGNCGSG 300 VEALPLQPNS EHEVLDQILE SSFTMQDFAS LQESMVKSQN QELNSGLTSD RTVWFQGQDM 360 ELNAISNLAS NEKAPYLSTM KQHLLHGALG EEGLKKMDSF NRWMSKELGD VGVIADANES 420 FTQSSSRTYW EEVESEDGSN GHNSRRDMDG YVMSPSLSKE QLFSINDFSP SWAYVGCEVV 480 VFVTGKFLKT REETEIGEWS CMFGQTEVPA DVISNGILQC VAPMHEAGRV PFYVTCSNRL 540 ACSEVREFEY KVAESQVFDR EADDESTIDI LEARFVKLLC SKSENTSPVS GNDSDLSQLS 600 EKISLLLFEN DDQLDQMLMN EISQENMKNN LLQEFLKESL HSWLLQKIAE GGKGPSVLDE 660 GGQGVLHFAA SLGYNWALEP TIIAGVSVDF RDVNGWTALH WAAFFGRERI IGSLIALGAA 720 PGTLTDPNPD FPSGSTPSDL AYANGHKGIA GYLSEYALRA HVSLLSLNDK NAETVEMAPS 780 PSSSSLTDSL TAVRNATQAA ARIHQVFRAQ SFQKKQLKEF GDKKLGMSEE RALSMLAPKT 840 HKSGRAHSDD SVQAAAIRIQ NKFRGYKGRK DYLITRQRII KIQAHVRGYQ FRKNYRKIIW 900 SVGVLEKVIL RWRRKGAGLR GFKSEALVEK MQDGTEKEED DDFFKQGRKQ TEDRLQKALA 960 RVKSMVQYPE ARDQYRRLLN VVNDIQESKV EKALENSEAT CFDDDDDLID IEALLEDDDT 1020 LMLPMSSSLW TS |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.48505 | 0.0 | flower| leaf| root | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 145361576 | 0.0 | ||||
| Genevisible | 263457_at | 0.0 | ||||
| Expression Atlas | AT2G22300 | - | ||||
| AtGenExpress | AT2G22300 | - | ||||
| ATTED-II | AT2G22300 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, carpels, and siliques, but not in stigmas or other parts of the flower. {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:14581622}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a putative CAM binding transcription factor. Loss of function mutations show enhanced resistance to fungal and bacterial pathogens suggesting that CAMTA functions to suppress defense responses. | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | 25215497 | Download |
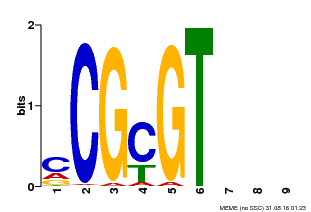
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT2G22300.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- ATRM (Manually Curated Upstream Regulators) ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator (A: Activate/R: Repress) | |||||
| ATRM | AT2G01570 (A) | |||||
| Regulation -- ATRM (Manually Curated Target Genes) ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Target Gene (A: Activate/R: Repress) | |||||
| ATRM | AT3G48090(R), AT4G25470(A) | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT2G22300 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF506697 | 0.0 | AF506697.1 Arabidopsis thaliana calmodulin-binding transcription factor SR1 (SR1) mRNA, complete cds. | |||
| GenBank | AY510025 | 0.0 | AY510025.1 Arabidopsis thaliana ethylene-induced calmodulin-binding protein 1 mRNA, complete cds. | |||
| GenBank | BT002459 | 0.0 | BT002459.1 Arabidopsis thaliana Unknown protein mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001118361.1 | 0.0 | signal responsive 1 | ||||
| Refseq | NP_850023.1 | 0.0 | signal responsive 1 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A178VUE5 | 0.0 | A0A178VUE5_ARATH; SR1 | ||||
| STRING | AT2G22300.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM8601 | 25 | 35 | Representative plant | OGRP562 | 15 | 65 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT2G22300.1 |
| Entrez Gene | 816762 |
| iHOP | AT2G22300 |
| wikigenes | AT2G22300 |



