 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vradi07g18170.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 296aa MW: 32234.8 Da PI: 7.5833 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 23.3 | 1.5e-07 | 8 | 53 | 4 | 45 |
S-HHHHHHHHHHHHHTTTT.....-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 4 WTteEdellvdavkqlGgg.....tWktIartmgkgRtlkqcksrwq 45
W eE++ + +a++++ + +W++Ia+ ++ ++ +++k ++q
Vradi07g18170.1 8 WNYEEEKAFENAIAMHWIEeasneQWEKIASAVP-SKSMEEVKQHYQ 53
999**************99***************.***********9 PP
| |||||||
| 2 | Myb_DNA-binding | 39 | 1.9e-12 | 135 | 179 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + ++G+g+W++I+r + Rt+ q+ s+ qky
Vradi07g18170.1 135 PWTEEEHRLFLLGLDKFGKGDWRSISRNFVISRTPTQVASHAQKY 179
7*****************************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 8.209 | 3 | 59 | IPR017884 | SANT domain |
| SMART | SM00717 | 2.5E-4 | 4 | 58 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-4 | 6 | 55 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 7.36E-7 | 8 | 53 | No hit | No description |
| SuperFamily | SSF46689 | 4.59E-9 | 8 | 62 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.873 | 128 | 184 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.11E-17 | 130 | 185 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.1E-17 | 132 | 182 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.3E-10 | 132 | 182 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.3E-11 | 133 | 178 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 9.64E-10 | 135 | 180 | No hit | No description |
| Pfam | PF00249 | 7.2E-11 | 135 | 179 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 296 aa Download sequence Send to blast |
MSSSGSLWNY EEEKAFENAI AMHWIEEASN EQWEKIASAV PSKSMEEVKQ HYQILVEDVS 60 AIEAGHIPFP NYTSEETTSS NKDFHGSSKA TSSDKRSNCN YSTGFSGLGH DSTTHSTGKG 120 ALSRSSEQER RKGIPWTEEE HRLFLLGLDK FGKGDWRSIS RNFVISRTPT QVASHAQKYF 180 IRLNSMNRDR RRSSIHDITS VNNGDVASNQ APITGQHSST VSSSTMGVGQ SLKHRVQGHI 240 APGLGMYGTP VGHPVAAPPG HMASAVGTPV MLPPGPHPPY VLPLAYPMAP PTMHQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 5e-15 | 4 | 74 | 7 | 75 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that coordinates abscisic acid (ABA) biosynthesis and signaling-related genes via binding to the specific promoter motif 5'-(A/T)AACCAT-3'. Represses ABA-mediated salt (e.g. NaCl and KCl) stress tolerance. Regulates leaf shape and promotes vegetative growth. {ECO:0000269|PubMed:26243618}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00191 | DAP | Transfer from AT1G49010 | Download |
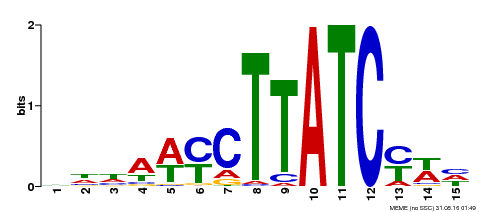
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vradi07g18170.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salicylic acid (SA) and gibberellic acid (GA) (PubMed:16463103). Triggered by dehydration and salt stress (PubMed:26243618). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:26243618}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KT031203 | 0.0 | KT031203.1 Glycine max clone HN_CCL_93 MYB/HD-like transcription factor (Glyma17g13010.1) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014507110.1 | 0.0 | transcription factor MYBS1 | ||||
| Swissprot | Q9FNN6 | 5e-86 | SRM1_ARATH; Transcription factor SRM1 | ||||
| TrEMBL | A0A1S3UM88 | 0.0 | A0A1S3UM88_VIGRR; transcription factor MYBS1 | ||||
| STRING | XP_007155139.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF10694 | 33 | 40 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G49010.1 | 5e-78 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



