 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT1G49010.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 314aa MW: 34471.7 Da PI: 9.323 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 23 | 1.9e-07 | 7 | 54 | 3 | 45 |
SS-HHHHHHHHHHHHHTTTT......-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg......tWktIartmgkgRtlkqcksrwq 45
W+ eE++ + +a++++ + +W++ +++++ + l+++k ++q
AT1G49010.1 7 TWSREEEKAFENAIALHCVEeeitedQWNKMSSMVP-SKALEEVKKHYQ 54
5****************999****************.***********9 PP
| |||||||
| 2 | Myb_DNA-binding | 38.8 | 2.1e-12 | 135 | 179 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + ++G+g+W++I+r + Rt+ q+ s+ qky
AT1G49010.1 135 PWTEEEHRLFLLGLDKFGKGDWRSISRNFVISRTPTQVASHAQKY 179
7*****************************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 6.975 | 3 | 61 | IPR017884 | SANT domain |
| SMART | SM00717 | 0.0017 | 4 | 59 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.13E-6 | 7 | 57 | No hit | No description |
| SuperFamily | SSF46689 | 2.07E-8 | 8 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.873 | 128 | 184 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.59E-17 | 130 | 185 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.5E-17 | 132 | 182 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.3E-10 | 132 | 182 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.9E-11 | 133 | 178 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.13E-9 | 135 | 180 | No hit | No description |
| Pfam | PF00249 | 7.9E-11 | 135 | 179 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 314 aa Download sequence Send to blast |
MESVVATWSR EEEKAFENAI ALHCVEEEIT EDQWNKMSSM VPSKALEEVK KHYQILLEDV 60 KAIENGQVPL PRYHHRKGLI VDEAAAAATS PANRDSHSSG SSEKKPNPGT SGISSSNGGR 120 SGGSRAEQER RKGIPWTEEE HRLFLLGLDK FGKGDWRSIS RNFVISRTPT QVASHAQKYF 180 IRLNSMNRDR RRSSIHDITT VNNQAPAVTG GGQQPQVVKH RPAQPQPQPQ PQPQQHHPPT 240 MAGLGMYGGA PVGQPIIAPP DHMGSAVGTP VMLPPPMGTH HHHHHHHLGV APYAVPAYPV 300 PPLPQQHPAP STMH |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 5e-13 | 8 | 78 | 11 | 78 | RADIALIS |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 30694467 | 0.0 | ||||
| Genevisible | 260769_at | 0.0 | ||||
| Expression Atlas | AT1G49010 | - | ||||
| AtGenExpress | AT1G49010 | - | ||||
| ATTED-II | AT1G49010 | - | ||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
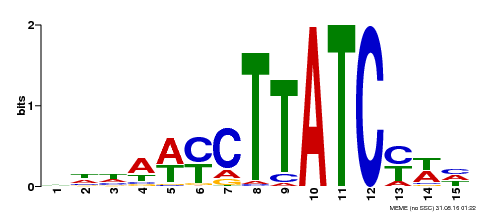
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00191 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT1G49010.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- Hormone ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hormone | |||||
| AHD | auxin, gibberellin, jasmonic acid, salicylic acid | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT1G49010 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY086906 | 0.0 | AY086906.1 Arabidopsis thaliana clone 29302 mRNA, complete sequence. | |||
| GenBank | AY519528 | 0.0 | AY519528.1 Arabidopsis thaliana MYB transcription factor (At1g49010) mRNA, complete cds. | |||
| GenBank | BT024862 | 0.0 | BT024862.1 Arabidopsis thaliana At1g49010 gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_564537.1 | 0.0 | Duplicated homeodomain-like superfamily protein | ||||
| TrEMBL | Q9M9A3 | 0.0 | Q9M9A3_ARATH; At1g49010 | ||||
| STRING | AT1G49010.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM10304 | 25 | 35 | Representative plant | OGRP275 | 17 | 122 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT1G49010.1 |
| Entrez Gene | 841324 |
| iHOP | AT1G49010 |
| wikigenes | AT1G49010 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



