 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vradi02g12120.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 397aa MW: 45150.9 Da PI: 6.2073 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 88.2 | 9.7e-28 | 66 | 148 | 2 | 86 |
trihelix 2 WtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqle 86
W ++e++ Li +rrem+ ++++k++k+lWe++s+kmre+gf rsp++C++kw+nl k++kk k++++++ s +++y+++++
Vradi02g12120.1 66 WVQDETRSLIGLRREMDALFNTSKSNKHLWEQISAKMREKGFDRSPTMCTDKWRNLLKEFKKAKHQDRGG--SGSAKMSYYKEID 148
********************************************************************98..34558*****997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 5.2E-4 | 57 | 127 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50090 | 8.455 | 58 | 122 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.0011 | 62 | 124 | IPR001005 | SANT/Myb domain |
| CDD | cd12203 | 6.39E-29 | 64 | 129 | No hit | No description |
| Pfam | PF13837 | 7.8E-21 | 65 | 150 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 397 aa Download sequence Send to blast |
MYLSEKPRPI DFYKEEGARD MMIEVVSNGE LAPPPPHHPP PMILGESSGE DPEVEIKAPK 60 KRAETWVQDE TRSLIGLRRE MDALFNTSKS NKHLWEQISA KMREKGFDRS PTMCTDKWRN 120 LLKEFKKAKH QDRGGSGSAK MSYYKEIDEI LRERNKNVQY KSPTPPPKVD SFMQFADKGS 180 DHSRNLTFEG LKGVVSSCID DTSISFGPVE ATGRPTLNLE RSLDHDGHPL AITTADAVAA 240 GGVPPWNWRE TSGNGGESQS CCGRVISVKW CDYTRRIGID GTPEAIKEAI RAAFRLRTKR 300 TFWLEDEDQI IRSIDREMPI GSYTLHLDEG MAIKLCLYDE SEHLPVHTED KVFYTENDYR 360 DFLTRRGWVC LREFDGYRNI DNLDDLRPGA IYRGVS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2jmw_A | 4e-48 | 60 | 146 | 1 | 86 | DNA binding protein GT-1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
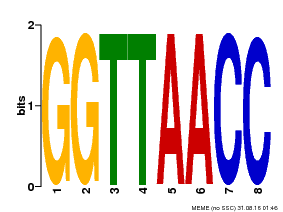
| UniProt | Probable transcription factor that binds specifically to the core DNA sequence 5'-GGTTAA-3'. May act as a molecular switch in response to light signals. {ECO:0000269|PubMed:10437822, ECO:0000269|PubMed:15044016, ECO:0000269|PubMed:7866025}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00004 | PBM | Transfer from 920224 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vradi02g12120.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015037 | 0.0 | AP015037.1 Vigna angularis var. angularis DNA, chromosome 4, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014494379.1 | 0.0 | trihelix transcription factor GT-1 isoform X1 | ||||
| Swissprot | Q9FX53 | 0.0 | TGT1_ARATH; Trihelix transcription factor GT-1 | ||||
| TrEMBL | A0A1S3TKS5 | 0.0 | A0A1S3TKS5_VIGRR; trihelix transcription factor GT-1 isoform X1 | ||||
| STRING | XP_007163722.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF6817 | 32 | 48 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G13450.1 | 0.0 | Trihelix family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



