 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vocar.0003s0248.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Chlorophyceae; Chlamydomonadales; Volvocaceae; Volvox
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 987aa MW: 100016 Da PI: 4.5885 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 53.8 | 2.6e-17 | 220 | 249 | 1 | 30 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyy 30
C+ Cg+t+Tp+WR+gp g+ktLCnaCG++y
Vocar.0003s0248.1.p 220 CVECGATQTPQWREGPAGPKTLCNACGVRY 249
****************************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd04370 | 2.04E-10 | 34 | 148 | No hit | No description |
| PROSITE profile | PS51038 | 16.558 | 35 | 159 | IPR001025 | Bromo adjacent homology (BAH) domain |
| SMART | SM00439 | 9.0E-8 | 35 | 159 | IPR001025 | Bromo adjacent homology (BAH) domain |
| Pfam | PF01426 | 7.6E-8 | 35 | 156 | IPR001025 | Bromo adjacent homology (BAH) domain |
| PROSITE profile | PS50114 | 12.479 | 214 | 249 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 1.8E-11 | 214 | 270 | IPR000679 | Zinc finger, GATA-type |
| Gene3D | G3DSA:3.30.50.10 | 4.7E-14 | 217 | 252 | IPR013088 | Zinc finger, NHR/GATA-type |
| SuperFamily | SSF57716 | 6.65E-12 | 217 | 254 | No hit | No description |
| CDD | cd00202 | 2.70E-12 | 219 | 269 | No hit | No description |
| Pfam | PF00320 | 1.0E-14 | 220 | 249 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 220 | 245 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 987 aa Download sequence Send to blast |
MSGPLQVEFF FSGPPLNANL GASRTDYAGF NLNGVEYKLG DCAYLYPEDD GMPPYVGRIL 60 ACFHDRSGRA ADPHCVEVAW YERRAHLEPD PHGGNWDREV VALEETDTNP IGCISGKAFV 120 FRAGDYEEAC RMAKSVAVDD WLFCRGVYKQ CENCFAPFPD LAEQMEVEDG RRHGKRSQAG 180 ISGHVEGVAD SEEVKRQRVA EEQPAAPVRR TGKLVSGRTC VECGATQTPQ WREGPAGPKT 240 LCNACGVRYV RAQQRANKRA VPAGAARSGY GAGGRQSRTS KAAKAEVASA RAARTAAVVP 300 AAAAGMAPGE EASQRPQRQA ALMAASRTAQ YARTGFFPLD TIDLQQQQQQ HTPDNAAING 360 QIAMHHQHHQ HEADMGDVLT GAFGGVSSAG AAAAGAAAAG PGSPNCSSYD SCGSLQEGVT 420 VEVVVGAADG EAECCGGSVA AVAPAAAAAA TAATVPTYPL LTSSSVGAAA GSLMPTPAQL 480 IGGSGIMLGG VEESVMAVAP VMALPLGEGQ LQSAVPGALQ TASAVAAAVA AAGVLGTASS 540 EFVLECDTTG SGMFFASGLD CQFSGDGVCY PADSHHHHHH HHDDHDQNGV CHHSKTGCVA 600 PAPSGGQCDT AAVYQGTGAV SCVAQVKSVP VSPCVDVDGA GFGCGGFGSV SRMLDGVLAS 660 ARGYGSQLGC GLVGGLDDEY TPLVEVEVDF GLGPASAMGD GSLFDGFDSG TCGADTVEDD 720 DPQHPQSQPL LAQGPALSAS QSQLQLPLPT TLVQQQRQLQ QQHEEQGDYG EGNMGVVDLS 780 AVAARVAAFC PDLAPGTDAR TLLDSLPLLA QTAFLPHGAG SAAAAALAVG DADAISLVAA 840 AAPTSVTVSA ADSAASDEGS CSVGVAVGGA VPGEAVKHLV ALGRQAQCAA TEAMAADAAL 900 QAANQVLEAH QEAARKAGEV AVSKLGLLKD CISTLAGGGE PLVAGEVGGH FIDLHDLGTV 960 GGLGDMAATG CCYSFDAADL LVAVHS* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00048 | PBM | Transfer from AT4G32890 | Download |
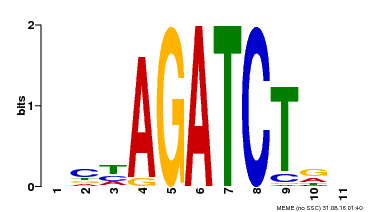
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002957265.1 | 0.0 | hypothetical protein VOLCADRAFT_98336 | ||||
| TrEMBL | D8UF31 | 0.0 | D8UF31_VOLCA; Uncharacterized protein | ||||
| STRING | XP_002957265.1 | 0.0 | (Volvox carteri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP220 | 14 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G32890.1 | 8e-15 | GATA transcription factor 9 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Vocar.0003s0248.1.p |



