 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vang0002ss01450.2 | ||||||||
| Common Name | LR48_Vigan06g068800 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 270aa MW: 30339.1 Da PI: 9.1959 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 104.5 | 6.4e-33 | 21 | 75 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kprlrWt +LH+rFv+av++LGG++kAtPk++l+lm++kgLtl+h+kSHLQkYRl
Vang0002ss01450.2 21 KPRLRWTADLHDRFVDAVTKLGGPDKATPKSVLRLMGLKGLTLYHLKSHLQKYRL 75
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 6.0E-32 | 18 | 76 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 12.455 | 18 | 78 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.69E-16 | 21 | 78 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.7E-24 | 21 | 76 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.8E-9 | 23 | 74 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 2.5E-23 | 119 | 165 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 270 aa Download sequence Send to blast |
MEGGGRESYN GLVMTMTRDP KPRLRWTADL HDRFVDAVTK LGGPDKATPK SVLRLMGLKG 60 LTLYHLKSHL QKYRLGQQTR KQNEEMNKEN SRCSFVNFST RSSGPNTTYR GDNEGGETPI 120 AEALRHQIEV QKRLEEQLEV QKKLQMRIEA QGKYLQAVLE KAQRTLSLDG SGGLEASRAQ 180 LTEFNSALSN FMENMNKDSK ENIIEVTGFY NKNHGSAFHY EEVGREQNRD QKPKIDGGSI 240 QFDLNIKGSN DLMSSGGAEM DVNMISYRG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-19 | 21 | 74 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-19 | 21 | 74 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-19 | 21 | 74 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-19 | 21 | 74 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-19 | 21 | 74 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-19 | 21 | 74 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-19 | 21 | 74 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-19 | 21 | 74 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
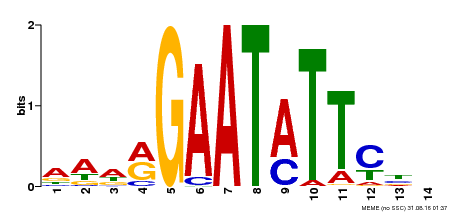
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00544 | DAP | Transfer from AT5G45580 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vang0002ss01450.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015043 | 0.0 | AP015043.1 Vigna angularis var. angularis DNA, chromosome 10, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017426744.1 | 0.0 | PREDICTED: myb family transcription factor PHL11 | ||||
| Swissprot | C0SVS4 | 4e-81 | PHLB_ARATH; Myb family transcription factor PHL11 | ||||
| TrEMBL | A0A0L9US08 | 0.0 | A0A0L9US08_PHAAN; Uncharacterized protein | ||||
| TrEMBL | A0A0S3T409 | 0.0 | A0A0S3T409_PHAAN; Uncharacterized protein | ||||
| STRING | XP_007158199.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G45580.1 | 1e-82 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



