 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRIUR3_17409-P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 361aa MW: 39443.9 Da PI: 4.7811 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 63.2 | 5.4e-20 | 104 | 154 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
+g++GVr++ +g+WvAeIr+p + r r +lg+f tae+Aa+a+++a+++++g
TRIUR3_17409-P1 104 CGFRGVRQRT-WGKWVAEIREP---N-RvSRLWLGTFPTAEDAARAYDEAARAMYG 154
79*****999.**********8...2.58************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.63E-30 | 104 | 164 | No hit | No description |
| Pfam | PF00847 | 3.3E-13 | 104 | 154 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 4.71E-21 | 105 | 163 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 22.3 | 105 | 162 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 7.2E-35 | 105 | 168 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.7E-10 | 106 | 117 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.4E-31 | 106 | 163 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.7E-10 | 128 | 144 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0010286 | Biological Process | heat acclimation | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 361 aa Download sequence Send to blast |
MGTSRLTLEK SAGSSNLPCN EYAFLARQNP KGDALPVASI LRKKRPRRSR DGPNSVSETI 60 RRWKEVNQQL EHDPQGAKRA RKPPAKGSKK GCMQGKGGPE NTQCGFRGVR QRTWGKWVAE 120 IREPNRVSRL WLGTFPTAED AARAYDEAAR AMYGALARTN FPAHPAQAPA VALPAAIEGV 180 VRGASASCES TTTSANHSDV ASNLPRQAQA LEIYSQPDVL ESTESVVLES VEHYSHKDTV 240 PDAGSSIARS TSEEDVFEPL EPISSLPDGE SDGFDIEELL RLMEADPIEV EPVTGGSWNC 300 GTNTGVEMGL QEPLYLDGLD QGMLEGMLQA DYPYPMWISE DRAMHNPAFH DAEMSEFFEG 360 L |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 6e-15 | 106 | 162 | 6 | 63 | ATERF1 |
| 3gcc_A | 6e-15 | 106 | 162 | 6 | 63 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3' of the cis-acting dehydration-responsive element (DRE). Binding to the C-repeat/DRE element mediates high salinity- and dehydration-inducible transcription. Involved in drought and heat-shock stress tolerance. {ECO:0000269|PubMed:20049613}. | |||||
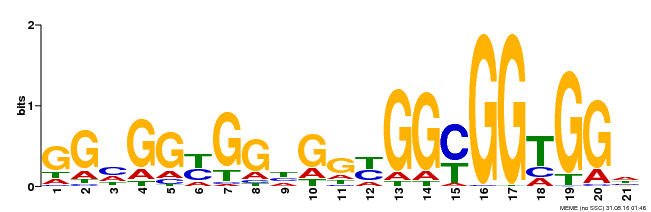
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00302 | DAP | Transfer from AT2G40340 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, high-salt and drought stresses. {ECO:0000269|PubMed:20049613}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY781354 | 0.0 | AY781354.1 Triticum aestivum DREB transcription factor 4A (DREB4) mRNA, complete cds, alternatively spliced. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020156297.1 | 0.0 | dehydration-responsive element-binding protein 2B-like isoform X1 | ||||
| Swissprot | Q5W6R4 | 3e-83 | DRE2B_ORYSJ; Dehydration-responsive element-binding protein 2B | ||||
| TrEMBL | M7YEC4 | 0.0 | M7YEC4_TRIUA; Dehydration-responsive element-binding protein 2B | ||||
| STRING | TRIUR3_17409-P1 | 0.0 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP11515 | 30 | 37 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40340.1 | 2e-36 | ERF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



