 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp57577_TGAC_v2_mRNA3907 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Trifolium
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 293aa MW: 32167.9 Da PI: 6.2419 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 51 | 2.5e-16 | 126 | 172 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn E+rRR++iN+++ L+ l+P++ K +Ka++L +A+eY+k+Lq
Tp57577_TGAC_v2_mRNA3907 126 FHNLSEKRRRSKINEKLKALQNLIPNS-----NKTDKASMLDEAIEYLKQLQ 172
5*************************7.....5******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 4.06E-20 | 119 | 182 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 5.2E-20 | 119 | 181 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.815 | 122 | 171 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.81E-18 | 125 | 176 | No hit | No description |
| Pfam | PF00010 | 8.1E-14 | 126 | 172 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.5E-18 | 128 | 177 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010114 | Biological Process | response to red light | ||||
| GO:0010154 | Biological Process | fruit development | ||||
| GO:0010187 | Biological Process | negative regulation of seed germination | ||||
| GO:0048440 | Biological Process | carpel development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 293 aa Download sequence Send to blast |
MLTAYQKTEN RTSSPFSLSL SLSLSLSLIM AGFSSDDIMH QTHPSLVPPE SQDFSNLLFN 60 QLMDPSSSSS SFNFSDPFPN PSTTIPYPPS HFTSSSTQND EASELPSSKA TPPPRSSSKR 120 SRAAEFHNLS EKRRRSKINE KLKALQNLIP NSNKTDKASM LDEAIEYLKQ LQLQVQMLMV 180 RNGYSLHPMS LSGGPRPTMY SQTDLLNLDE GNVGFHNSIN ANDECYARPG FSFPEQCSIS 240 NQSVILPSAT NIANFDASSS FQPSIKDVFC NNMPQLILDS TRIGKTPSPD VS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Required for the dehiscence of fruit, especially for the separation of the valve cells from the replum. Promotes the differentiation of a strip of labile nonlignified cells sandwiched between layers of lignified cells. {ECO:0000269|PubMed:11747817}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00035 | PBM | Transfer from AT4G36930 | Download |
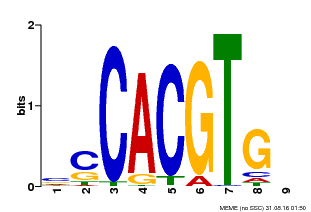
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp57577_TGAC_v2_mRNA3907 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT147727 | 1e-82 | BT147727.1 Lotus japonicus clone JCVI-FLLj-8F4 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003589168.2 | 1e-143 | transcription factor ALC | ||||
| Swissprot | Q9FHA2 | 8e-35 | ALC_ARATH; Transcription factor ALC | ||||
| TrEMBL | A0A2Z6NSW4 | 1e-175 | A0A2Z6NSW4_TRISU; Uncharacterized protein | ||||
| STRING | AES59419 | 1e-127 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2276 | 33 | 79 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G67110.1 | 1e-24 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Tp57577_TGAC_v2_mRNA3907 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



