 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp4g28550 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 573aa MW: 63718.5 Da PI: 5.8567 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 43.7 | 5e-14 | 400 | 446 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr+++P+ + K++Ka+ L A+ YIk+Lq
Tp4g28550 400 NHVEAERQRREKLNQRFYALRSVVPNiS------KMDKASLLGDAISYIKELQ 446
799***********************66......******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.4E-50 | 48 | 239 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| SuperFamily | SSF47459 | 4.58E-19 | 395 | 460 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.936 | 396 | 445 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.45E-15 | 399 | 450 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 1.1E-18 | 400 | 460 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.9E-11 | 400 | 446 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 6.3E-18 | 402 | 451 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 573 aa Download sequence Send to blast |
MNMSDLGWDD EEKALVSAVI GNLASDFLRA NSNSNQNLFL VMGTDDNLNK KLSSLVDWQN 60 SENFSWNYAI FWQKTMSRSG HQVLGWGDGC CREPNVEEES KVVRVYNCNS LGLEEQRCQD 120 MRKTVLQKLH RLFGGSDEDN YALSLENVTP TEIFFLASMY FFFNHGEGGP GRCYASGKHV 180 WLSDAVNSVS DYCFRSFMAK SAGIRTIVMV PTDAGVLELG SVWSLPENVE LVKSVQALFM 240 RRAKLPLMVN STNTNMSGGG IHKLFGQELN NSEQGAHHGK KLDVRNLDQR FIPQIWQGYN 300 NNKDSTFGYT SQGIDVKVQE NENVVVDDNN YKMQIEFAGS SVAASSNPST NTQLEKSDSC 360 TEKRPASLLA GVAGTVSLVD EKRPRKRGRK PANGREEPLN HVEAERQRRE KLNQRFYALR 420 SVVPNISKMD KASLLGDAIS YIKELQEKVK IMEADRVRTE SSLAESNPMT TRTVESPEVD 480 IHAVNEEVVV RVISPLDSHP ASSIIQAMRN SEVSLTESKL SLAEDTMFHT FVIKSNNGSD 540 PLTKEKLIAA AYPQTSSTQQ PLPASSSQVS GDI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 8e-29 | 394 | 465 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_B | 8e-29 | 394 | 465 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_E | 8e-29 | 394 | 465 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_F | 8e-29 | 394 | 465 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_G | 8e-29 | 394 | 465 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_I | 8e-29 | 394 | 465 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_M | 8e-29 | 394 | 465 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_N | 8e-29 | 394 | 465 | 2 | 73 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 381 | 389 | KRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator. Regulates positively abscisic acid (ABA) response. Confers drought tolerance and sensitivity to ABA. {ECO:0000269|PubMed:17828375}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00101 | PBM | Transfer from AT2G46510 | Download |
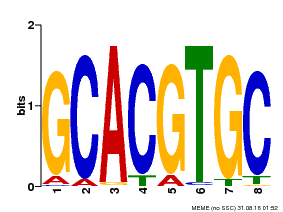
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp4g28550 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Transiently by ABA and polyethylene glycol (PEG). {ECO:0000269|PubMed:17828375}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006397843.1 | 0.0 | transcription factor ABA-INDUCIBLE bHLH-TYPE | ||||
| Swissprot | Q9ZPY8 | 0.0 | AIB_ARATH; Transcription factor ABA-INDUCIBLE bHLH-TYPE | ||||
| TrEMBL | V4N299 | 0.0 | V4N299_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006397843.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3758 | 27 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46510.1 | 0.0 | ABA-inducible BHLH-type transcription factor | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



