 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp4g01350 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1025aa MW: 115078 Da PI: 5.0201 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 182.2 | 5.8e-57 | 20 | 136 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywl 101
+e ++rwl++ ei++iL+n++++++++e++t+p sgs++L++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+v+vl+cyYah+++n++fqrr+yw+
Tp4g01350 20 SEaRNRWLRPPEICEILQNHQRFQISTEPPTTPASGSVFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKAGSVDVLHCYYAHGQDNENFQRRSYWM 119
4449************************************************************************************************ PP
CG-1 102 Leeelekivlvhylevk 118
L+eel++iv+vhylevk
Tp4g01350 120 LQEELSHIVFVHYLEVK 136
**************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 83.413 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.6E-84 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 6.7E-50 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 7.52E-15 | 451 | 536 | IPR014756 | Immunoglobulin E-set |
| CDD | cd00204 | 5.06E-11 | 613 | 743 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 7.1E-15 | 614 | 747 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 16.733 | 617 | 743 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 3.58E-15 | 629 | 746 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.683 | 684 | 716 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 2.31E-8 | 842 | 896 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.4 | 845 | 867 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.139 | 847 | 875 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.002 | 848 | 866 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0019 | 868 | 890 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.56 | 869 | 893 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 1.6E-4 | 871 | 890 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1025 aa Download sequence Send to blast |
MAEARRFSPN NELDVGQILS EARNRWLRPP EICEILQNHQ RFQISTEPPT TPASGSVFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHER LKAGSVDVLH CYYAHGQDNE NFQRRSYWML 120 QEELSHIVFV HYLEVKGSRV STSNNRMQRT EDSARSSQET GELFTSERDG YASASINQYD 180 HSNHSQATDS ASVNGVHTPE LEDAESAYNQ QGSSIVYSHQ ELQQPATTGF DPYYQMTLTP 240 RDSYQKELRT VPITDSSIMV DKSRTINGPG VTNGLRNKKS IDSQTWEEIL GNCGSGAEGL 300 PVQPNSENEV LDQILQSYSF TMQDFASLQE SMVKSQNQEL NSVLTSDRTM WFQGQDLELN 360 AISNLASNEK APYLSTMKQH LLDGALGEEE LGELGDVGVI ADANESFTHS SSTAMWEEVE 420 SEDRSNGHNS RRDLDGYVIS PSLSKEQLFS IIDFAPSWAY VGFEVKVLVT GKFLKTREEA 480 EIGEWSCMFG QTEVPADVIA NGVLQCVAPM LEAGRVPFYV TCSNRLACSE VREFEYKVAE 540 SQVFDREKDE SSACNSIESF EARFLKLLCS KSDCLSSSPS ANDSDLSELS EKISLLLFEN 600 DDQLDQMLMN EISQENMKNN LLQEALKESL HSWLLQKIAE GGKGPNVLDE GGQGILHFAA 660 ALGYNWALEP TIVAGVSVDF RDINGWTALH WAAFFGRELI IGSLIALGAA PGTLTDPNPD 720 FPSGSTPSDL AYANGYKGIA GYLSEYALRA HVSLLSLNEK NAETSLAAVE TTPSQSSSSL 780 ADSLTAVRNA TQAAARIHQV FRAQSFQKKQ IKEFGDRKYG MSEERALSML APKTHKPGRA 840 HIDDSVQAAA IRIQNKFRGY KGRKEYLITR QRIIKIQAHV RGHQVRKNYR KIIWSVGILE 900 KVILRWRRKG AGLRGFKSDS LVDKMQDGTQ KEEDDDFFKQ GRKQTEERLE KALARVKSMV 960 QYPEARDQYR RLLNVVNDIQ ESKVEKALEN SEATCFDDDL IDIEALLEDD DDTLMLPMSS 1020 TMWNT |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
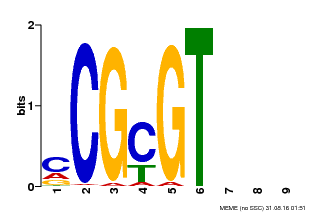
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp4g01350 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF506697 | 0.0 | AF506697.1 Arabidopsis thaliana calmodulin-binding transcription factor SR1 (SR1) mRNA, complete cds. | |||
| GenBank | AK226486 | 0.0 | AK226486.1 Arabidopsis thaliana mRNA for Calmodulin-binding transcription activator 3, complete cds, clone: RAFL06-74-H09. | |||
| GenBank | AY510025 | 0.0 | AY510025.1 Arabidopsis thaliana ethylene-induced calmodulin-binding protein 1 mRNA, complete cds. | |||
| GenBank | BT002459 | 0.0 | BT002459.1 Arabidopsis thaliana Unknown protein mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024014331.1 | 0.0 | calmodulin-binding transcription activator 3 isoform X2 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | V4LQS4 | 0.0 | V4LQS4_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006404687.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM8601 | 25 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



