 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp1g01020 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 590aa MW: 65703 Da PI: 7.0914 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.9 | 7.5e-13 | 433 | 478 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr+++P+ + K++Ka+ L Av YI++L
Tp1g01020 433 NHVEAERQRREKLNQRFYALRSVVPNiS------KMDKASLLGDAVSYINEL 478
799***********************66......***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 5.9E-50 | 48 | 236 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 17.326 | 429 | 478 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 8.9E-19 | 432 | 499 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 4.21E-14 | 432 | 483 | No hit | No description |
| Pfam | PF00010 | 2.0E-10 | 433 | 478 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 6.1E-18 | 433 | 492 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 4.8E-16 | 435 | 484 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 590 aa Download sequence Send to blast |
MNIGGVVWNE DDKAIVASLL GKRAIDYLLS SSVPNANLLM TVGSDENLQN KLSDLVERPN 60 ASNFSWNYAI FWQISRSKAG DLVLCWGDGS CREPKEGEKS EIVRILSMGR EEETHQTMRK 120 RVLQKLHALF GGLEEDNCAL RLDRVTDTEM FLLASMYFSF PRGEGGPGMC FDSGKPVWLP 180 DVVNSGSDYC VRSFLAKSAG IQTIVLVPTD IGVVELGSTR CLPESEESML SIRSSFSSNL 240 PPPVRVVTTA ALPVVVADKS DDNRTVTSSK IFGKDLHNSG FLRHHQTQQP QQHRQFREKL 300 TVRKMDDKVP KRVDAYPNNG NRYMFSNPST NNNTLLSPTW VQPEMYRRTN TVKEVPSTED 360 FKFLPLQQSS QRLPPPAQMQ IDFSGASSRA SENNSDGEGG AEWADVVGGD ESGNNRPRKR 420 GRRPANGRAE ALNHVEAERQ RREKLNQRFY ALRSVVPNIS KMDKASLLGD AVSYINELHA 480 KLKVMEAERE RLGYSSNPPI SLEPEVSVQT AGEDVTVRVN CPLGSHPASR IFHAFEESKV 540 EVISSQLEVP QDIVLHTFVI KSEELTKEKL ISAFSREQSS SVQSRTSSGR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 2e-26 | 427 | 512 | 2 | 80 | Transcription factor MYC2 |
| 5gnj_B | 2e-26 | 427 | 512 | 2 | 80 | Transcription factor MYC2 |
| 5gnj_E | 2e-26 | 427 | 512 | 2 | 80 | Transcription factor MYC2 |
| 5gnj_F | 2e-26 | 427 | 512 | 2 | 80 | Transcription factor MYC2 |
| 5gnj_G | 2e-26 | 427 | 512 | 2 | 80 | Transcription factor MYC2 |
| 5gnj_I | 2e-26 | 427 | 512 | 2 | 80 | Transcription factor MYC2 |
| 5gnj_M | 2e-26 | 427 | 512 | 2 | 80 | Transcription factor MYC2 |
| 5gnj_N | 2e-26 | 427 | 512 | 2 | 80 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
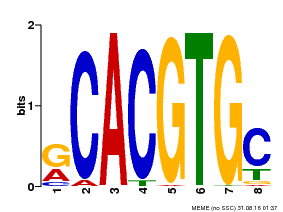
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp1g01020 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By UV treatment. {ECO:0000269|PubMed:12679534}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353331 | 0.0 | AK353331.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-24-P08. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006418406.1 | 0.0 | transcription factor bHLH13 | ||||
| Swissprot | Q9LNJ5 | 0.0 | BH013_ARATH; Transcription factor bHLH13 | ||||
| TrEMBL | E4MXT4 | 0.0 | E4MXT4_EUTHA; mRNA, clone: RTFL01-24-P08 | ||||
| TrEMBL | V4KZG7 | 0.0 | V4KZG7_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006418406.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3758 | 27 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 0.0 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



