 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010559123.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 592aa MW: 65919.7 Da PI: 5.7735 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 40 | 6.9e-13 | 419 | 465 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er RR+++N++f Lr+++P+ + K++Ka+ L A+ YI++Lq
XP_010559123.1 419 NHVEAERMRREKLNQRFYALRSVVPNiS------KMDKASLLGDAIAYINELQ 465
799***********************66......******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 6.9E-55 | 48 | 236 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| SuperFamily | SSF47459 | 2.75E-18 | 414 | 476 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 16.825 | 415 | 464 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.34E-14 | 418 | 469 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 5.8E-18 | 419 | 476 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 2.7E-10 | 419 | 465 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 9.9E-17 | 421 | 470 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 592 aa Download sequence Send to blast |
MNMGERGWDD EEKAMVSAVL GQRAFDYLRA SPVSNENLFA TMGSDESLQI KLSDLVDRPN 60 SMNFSWNYAI FWQLSQSRSG HLVLVWGDGC CREPNKEEES EAVGARNGGL EEDNWQNMRK 120 RVLQKLHRFS GGSDEDNYAL GLDRVTETEM FFLVSMYFSF PNGEGGPGRC FASGNHFWIS 180 DAMNTASDYC FRSFLAKSAG IRTIVMVPMD MGVVELGSVR SLPENMELVH SVRSLFSTRV 240 HQTATLFPYP MNERRDDDNN VEKNTGGVHK LFGQELGHSN SNHCHAFPPF RENMASRKMD 300 ERFPPQPWPE GYQSPRNGFH ALTPHAAQTT ALYAAQTTDS NMQGQVNATK EDAQLNNHQS 360 PKPTSQMQID YAASSKPSPN TVSRNLDSCL EKRAAAAMEE KRPRKRGRKP ANGREEPLNH 420 VEAERMRREK LNQRFYALRS VVPNISKMDK ASLLGDAIAY INELQEKVKV MEAERTGTGS 480 SPSPGEPNPS LESSNLIPDV EFQTLNDEVV VQVSSPLDTH PVSRIIQAVR NSQVSVLESK 540 LSLINDTVFH TFVIKSDGSE VLTKEKLMAA VCPEVGSEEP LPSSGSQVSG SL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 2e-28 | 413 | 475 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 2e-28 | 413 | 475 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 2e-28 | 413 | 475 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 2e-28 | 413 | 475 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 2e-28 | 413 | 475 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 2e-28 | 413 | 475 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 2e-28 | 413 | 475 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 2e-28 | 413 | 475 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 400 | 408 | KRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator. Regulates positively abscisic acid (ABA) response. Confers drought tolerance and sensitivity to ABA. {ECO:0000269|PubMed:17828375}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00101 | PBM | Transfer from AT2G46510 | Download |
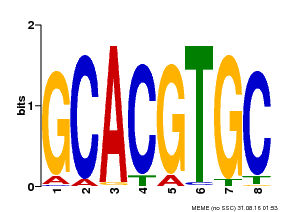
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010559123.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Transiently by ABA and polyethylene glycol (PEG). {ECO:0000269|PubMed:17828375}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010559123.1 | 0.0 | PREDICTED: transcription factor ABA-INDUCIBLE bHLH-TYPE-like | ||||
| Swissprot | Q9ZPY8 | 0.0 | AIB_ARATH; Transcription factor ABA-INDUCIBLE bHLH-TYPE | ||||
| TrEMBL | V4N299 | 0.0 | V4N299_EUTSA; Uncharacterized protein | ||||
| STRING | XP_010559123.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3758 | 27 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46510.1 | 0.0 | ABA-inducible BHLH-type transcription factor | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



