 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010535526.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 273aa MW: 31126.1 Da PI: 7.8801 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.5 | 2.7e-17 | 10 | 57 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT++Ed llv+ v+++G ++W+ Ia+ g++Rt+k+c++rw +yl
XP_010535526.1 10 KGPWTEQEDILLVNFVHLFGDRRWDFIAKVSGLNRTGKSCRLRWGNYL 57
79******************************************9996 PP
| |||||||
| 2 | Myb_DNA-binding | 41.7 | 2.7e-13 | 85 | 121 | 10 | 48 |
HHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 10 ellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
l ++++ ++G++ W++Iar+++ gRt++++k++w++++
XP_010535526.1 85 RLVLELHVKWGNR-WSKIARKLP-GRTDNEIKNYWRTHM 121
57789********.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 24.875 | 5 | 61 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.2E-15 | 9 | 59 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.91E-25 | 10 | 61 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 5.5E-16 | 10 | 57 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.8E-20 | 11 | 62 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.20E-11 | 12 | 57 | No hit | No description |
| PROSITE profile | PS51294 | 16.348 | 62 | 125 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.1E-8 | 62 | 123 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.7E-20 | 63 | 124 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 7.28E-9 | 84 | 121 | No hit | No description |
| Pfam | PF00249 | 5.0E-11 | 84 | 120 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.91E-25 | 93 | 122 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 273 aa Download sequence Send to blast |
MKMVQEEIRK GPWTEQEDIL LVNFVHLFGD RRWDFIAKVS GLNRTGKSCR LRWGNYLHPG 60 LKRGKMTPQE ERXXXXXXXX XXXERLVLEL HVKWGNRWSK IARKLPGRTD NEIKNYWRTH 120 MRKKAQEKKR SMSRTSSSSN CCSSTATTAT TQDSLQSPDS GSGKASFYDT GGFDGKLNQE 180 VEDCEGGYSM DDIWKEIDQS EASILKPVID IYAEQSCYLN YPPLASPAWE SSLDSIWNME 240 ADESKMSFAI DQFPHYSGYV RSSSSSPSSS LAG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h88_C | 6e-25 | 2 | 125 | 50 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| 1h89_C | 6e-25 | 2 | 125 | 50 | 159 | MYB PROTO-ONCOGENE PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00600 | PBM | Transfer from AT5G59780 | Download |
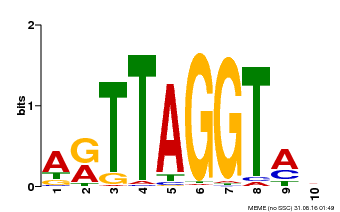
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010535526.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Isoform MYB59-1 is induced by jasmonate (JA), salicylic acid (SA), gibberellic acid (GA), and ethylene. Also induced by cadmium (Cd). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:16531467}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF062894 | 5e-57 | AF062894.1 Arabidopsis thaliana putative transcription factor (MYB59) mRNA, complete cds. | |||
| GenBank | AY519641 | 5e-57 | AY519641.1 Arabidopsis thaliana MYB transcription factor (At5g59780) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010535526.1 | 0.0 | PREDICTED: transcription factor MYB59 isoform X1 | ||||
| Swissprot | Q4JL84 | 1e-124 | MYB59_ARATH; Transcription factor MYB59 | ||||
| TrEMBL | A0A3P6ATB8 | 1e-123 | A0A3P6ATB8_BRAOL; Uncharacterized protein | ||||
| STRING | XP_010535526.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2502 | 27 | 74 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G59780.3 | 1e-119 | myb domain protein 59 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



