| Signature Domain? help Back to Top |
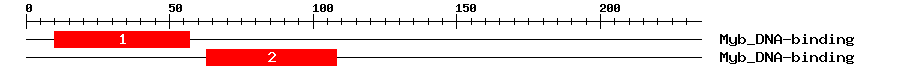 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Myb_DNA-binding | 55.4 | 1.5e-17 | 10 | 57 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT++Ed llv+ v+++G ++W+ +a+ g++Rt+k+c++rw +yl
AT5G59780.3 10 KGPWTEQEDILLVNFVHLFGDRRWDFVAKVSGLNRTGKSCRLRWVNYL 57
79********************************************97 PP
|
| 2 | Myb_DNA-binding | 56.4 | 7e-18 | 63 | 108 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T++E+ l +++++++G++ W++Iar+++ gRt++++k++w++++
AT5G59780.3 63 RGKMTPQEERLVLELHAKWGNR-WSKIARKLP-GRTDNEIKNYWRTHM 108
89********************.*********.************986 PP
|
| Publications
? help Back to Top |
- Riechmann JL, et al.
Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes.
Science, 2000. 290(5499): p. 2105-10
[PMID:11118137] - Stracke R,Werber M,Weisshaar B
The R2R3-MYB gene family in Arabidopsis thaliana.
Curr. Opin. Plant Biol., 2001. 4(5): p. 447-56
[PMID:11597504] - Nikiforova V, et al.
Transcriptome analysis of sulfur depletion in Arabidopsis thaliana: interlacing of biosynthetic pathways provides response specificity.
Plant J., 2003. 33(4): p. 633-50
[PMID:12609038] - Jiao Y, et al.
A genome-wide analysis of blue-light regulation of Arabidopsis transcription factor gene expression during seedling development.
Plant Physiol., 2003. 133(4): p. 1480-93
[PMID:14605227] - Contento AL,Kim SJ,Bassham DC
Transcriptome profiling of the response of Arabidopsis suspension culture cells to Suc starvation.
Plant Physiol., 2004. 135(4): p. 2330-47
[PMID:15310832] - Stanley Kim H, et al.
Transcriptional divergence of the duplicated oxidative stress-responsive genes in the Arabidopsis genome.
Plant J., 2005. 41(2): p. 212-20
[PMID:15634198] - Duarte JM, et al.
Expression pattern shifts following duplication indicative of subfunctionalization and neofunctionalization in regulatory genes of Arabidopsis.
Mol. Biol. Evol., 2006. 23(2): p. 469-78
[PMID:16280546] - He XJ, et al.
AtNAC2, a transcription factor downstream of ethylene and auxin signaling pathways, is involved in salt stress response and lateral root development.
Plant J., 2005. 44(6): p. 903-16
[PMID:16359384] - Yanhui C, et al.
The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family.
Plant Mol. Biol., 2006. 60(1): p. 107-24
[PMID:16463103] - Li J, et al.
A subgroup of MYB transcription factor genes undergoes highly conserved alternative splicing in Arabidopsis and rice.
J. Exp. Bot., 2006. 57(6): p. 1263-73
[PMID:16531467] - Cao D,Cheng H,Wu W,Soo HM,Peng J
Gibberellin mobilizes distinct DELLA-dependent transcriptomes to regulate seed germination and floral development in Arabidopsis.
Plant Physiol., 2006. 142(2): p. 509-25
[PMID:16920880] - Lee J, et al.
Analysis of transcription factor HY5 genomic binding sites revealed its hierarchical role in light regulation of development.
Plant Cell, 2007. 19(3): p. 731-49
[PMID:17337630] - Libault M,Wan J,Czechowski T,Udvardi M,Stacey G
Identification of 118 Arabidopsis transcription factor and 30 ubiquitin-ligase genes responding to chitin, a plant-defense elicitor.
Mol. Plant Microbe Interact., 2007. 20(8): p. 900-11
[PMID:17722694] - Ascencio-Ib
Global analysis of Arabidopsis gene expression uncovers a complex array of changes impacting pathogen response and cell cycle during geminivirus infection.
Plant Physiol., 2008. 148(1): p. 436-54
[PMID:18650403] - Agudelo-Romero P, et al.
Changes in the gene expression profile of Arabidopsis thaliana after infection with Tobacco etch virus.
Virol. J., 2008. 5: p. 92
[PMID:18684336] - Mu RL, et al.
An R2R3-type transcription factor gene AtMYB59 regulates root growth and cell cycle progression in Arabidopsis.
Cell Res., 2009. 19(11): p. 1291-304
[PMID:19581938] - Cominelli E,Tonelli C
A new role for plant R2R3-MYB transcription factors in cell cycle regulation.
Cell Res., 2009. 19(11): p. 1231-2
[PMID:19881525] - Ding Y, et al.
Four distinct types of dehydration stress memory genes in Arabidopsis thaliana.
BMC Plant Biol., 2013. 13: p. 229
[PMID:24377444] - Nishida S,Kakei Y,Shimada Y,Fujiwara T
Genome-wide analysis of specific alterations in transcript structure and accumulation caused by nutrient deficiencies in Arabidopsis thaliana.
Plant J., 2017. 91(4): p. 741-753
[PMID:28586097] - Hickman R, et al.
Architecture and Dynamics of the Jasmonic Acid Gene Regulatory Network.
Plant Cell, 2017. 29(9): p. 2086-2105
[PMID:28827376] - Romero I, et al.
More than 80R2R3-MYB regulatory genes in the genome of Arabidopsis thaliana.
Plant J., 1998. 14(3): p. 273-84
[PMID:9628022] - Kranz HD, et al.
Towards functional characterisation of the members of the R2R3-MYB gene family from Arabidopsis thaliana.
Plant J., 1998. 16(2): p. 263-76
[PMID:9839469]
|





