 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010525244.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 718aa MW: 78094.8 Da PI: 6.632 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.6 | 1e-16 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE+ ++++a k++G Wk+I +++g +t+ q++s+ qk+
XP_010525244.1 24 RERWTEEEHNRFLEALKLYGRA-WKKIEEHVG-SKTAVQIRSHAQKF 68
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.07E-17 | 18 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.997 | 19 | 73 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.8E-17 | 22 | 71 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.9E-13 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.7E-13 | 24 | 67 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.1E-9 | 24 | 64 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 5.62E-10 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010243 | Biological Process | response to organonitrogen compound | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043496 | Biological Process | regulation of protein homodimerization activity | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 718 aa Download sequence Send to blast |
METNSSGEEL VIKTRKPYTI TKERERWTEE EHNRFLEALK LYGRAWKKIE EHVGSKTAVQ 60 IRSHAQKFFS KVEKEAQAKG FPLGQALDVD IPPPRPKKKP SNPYPRKTGN GNPSQSAAKD 120 EKLPTSTSAL NCIEALVSEN GSLSEMPNEE RQPSNPEENR EDGCSECFTH SRSAYSMNKS 180 SVETTALVNG CTFREFLPLR KEASQDNGKI KASNSENYCD AHTKSVQGSV KNNVHGISHI 240 DDQGPQTYPR HVHTHGLDRS SLTCSMSPFP TVPDEIHGYP NMFVDPSASA TRSEHENDSP 300 RSTPRASPAF HPADAPTPNA PDNYRPFANL ILSTLLQNPA VHAAATFAAS VWPHNPGGPS 360 TCAEGGFPAG QVYSPPSLAA IAVPAATVAA ATAWWAAHGL LPLCGPFHSA FPCHPPPTNV 420 APSDDNGQIP LARVNPKENT LQKPSLQSRE EGPENFEASK AQFSPAKAVS SPSYSEDSER 480 RKTETGTKPG SNEQHAAETD QNGDKNTKQV VDRSSCGSNT SSSSDVEADA SEKLEKGNDE 540 VKEADADPNP PAAESNTRRS RSTNNLTDPW KSVSNEGRLA FQALFSRAVL PQSFSPPHDA 600 KKKKNRTENK DEETKKNTER ENGETLALDL NSKARAADSD HQEEEKGDGL GSGRENTDQK 660 GFLGIGLGTS KLKGRTGFKP YKRCSVEAKD NTIIDTTSLN QAEQKDSKRI RLESEAST |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the circadian clock and in the phytochrome regulation. Binds to the promoter regions of APRR1/TOC1 and TCP21/CHE to repress their transcription. Binds to the promoter regions of CAB2A and CAB2B to promote their transcription. Represses both LHY and itself. {ECO:0000269|PubMed:11486091, ECO:0000269|PubMed:12007421, ECO:0000269|PubMed:12015970, ECO:0000269|PubMed:19095940, ECO:0000269|PubMed:19218364, ECO:0000269|PubMed:19339503, ECO:0000269|PubMed:9657153}. | |||||
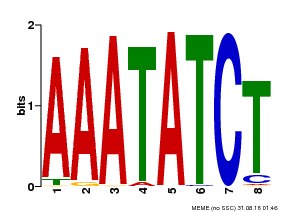
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00103 | PBM | Transfer from AT2G46830 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010525244.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation with peak levels occurring around 1 hour after dawn. Up-regulated by APRR1/TOC1 and transiently by light treatment. Down-regulated by APRR5, APRR7 and APRR9. The CCA1 mRNA is relatively stable in the dark and in far-red light but has a short half-life in red and blue light. {ECO:0000269|PubMed:17873091, ECO:0000269|PubMed:19095940, ECO:0000269|PubMed:19218364, ECO:0000269|PubMed:19286557, ECO:0000269|PubMed:20233950, ECO:0000269|PubMed:9144958, ECO:0000269|PubMed:9657153}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019056755.1 | 0.0 | PREDICTED: protein CCA1 | ||||
| Swissprot | P92973 | 1e-166 | CCA1_ARATH; Protein CCA1 | ||||
| TrEMBL | A0A061DT31 | 0.0 | A0A061DT31_THECC; Homeodomain-like superfamily protein isoform 1 | ||||
| STRING | XP_010525244.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5576 | 26 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 1e-137 | circadian clock associated 1 | ||||



