 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG041605t1 | ||||||||
| Common Name | TCM_041605 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 710aa MW: 76760.1 Da PI: 5.6981 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 20.2 | 1.1e-06 | 279 | 299 | 5 | 26 |
--SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 5 lCrffartGtCkyGdrCkFaHg 26
+C+ f++ G C+ Gd+C +aHg
Thecc1EG041605t1 279 PCPEFRK-GSCRQGDNCEYAHG 299
8******.*************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF48403 | 4.35E-15 | 51 | 150 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 4.5E-8 | 71 | 144 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.029 | 75 | 105 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50297 | 16.573 | 75 | 150 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 9.6E-16 | 76 | 174 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.69E-14 | 77 | 173 | No hit | No description |
| SMART | SM00248 | 8.1E-5 | 110 | 142 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 13.384 | 110 | 145 | IPR002110 | Ankyrin repeat |
| PRINTS | PR01415 | 8.0E-5 | 111 | 126 | IPR002110 | Ankyrin repeat |
| PRINTS | PR01415 | 8.0E-5 | 129 | 143 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 4400 | 152 | 181 | IPR002110 | Ankyrin repeat |
| SMART | SM00356 | 5.0E-5 | 274 | 300 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 4.9E-4 | 278 | 301 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 2.7E-4 | 279 | 299 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 13.094 | 279 | 301 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 6.5E-4 | 307 | 337 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 20 | 309 | 332 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 6.221 | 309 | 333 | IPR000571 | Zinc finger, CCCH-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 710 aa Download sequence Send to blast |
MCSGSKRKPT QTGFVMEGES HKEDMLPQKF SVLLEFSASN DLIGFKSAIE EEGHDIDEPS 60 LWYGRRIGLK KMGFEERTPL LIASMFGSKD VVNYIIGSGR VDVNRACGSD GATALHCAAA 120 GGSFDSAEVV KILLDASADI NSVDANGNRP GDLITPACNS AFSLRKKMLE ALLRGSGSVG 180 EMEALPAQKG NEMEGQEQQD NSIARVLKDG TEKKEYPVDL TLPDIKNGIY GTDEFRMYTF 240 KVKPCSRAYS HDWTECPFVH PGENARRRDP RKYHYSCVPC PEFRKGSCRQ GDNCEYAHGI 300 FECWLHPAQY RTRLCKDETN CSRRVCFFAH KPEELRPLYA STGSAVPSPR SYAANGSSLD 360 MGSMSPLALG SPSVMIPPTS TPPLTPTGTS SPMGGAMWPN QLSIVPPTLQ LPGSRLKTAL 420 SARDMDLDME LLGLESHRRR QQQQLIDEIS GLSSPTSWNN PLSSASAFSA TGDRTGELNR 480 FGGVKPTNLE DIFGSLDTTI LPQLQGISLD GTAPQLQSPT GVQMRQNINQ QLRASYPTNL 540 ASSPVRASPS FGIDASSGPT AAAVLSSRSA AFAKRSQSFI ERTAVNRHAG FSSPTSSASA 600 MPSNLSDWGS PDGKLDWGIQ GEELSKLRKS ASFGFRSSGS NLANAAPSML PTADEPDVSW 660 VQSLVKDTPS AGQFGFDEEQ QQCHLNSGGA EMLPAWVEQL YMEQEQMVA* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00566 | DAP | Transfer from AT5G58620 | Download |
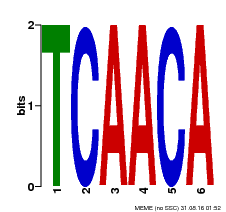
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007016084.2 | 0.0 | PREDICTED: zinc finger CCCH domain-containing protein 66 | ||||
| Swissprot | Q9LUZ4 | 0.0 | C3H66_ARATH; Zinc finger CCCH domain-containing protein 66 | ||||
| TrEMBL | A0A061GWD4 | 0.0 | A0A061GWD4_THECC; Zinc finger family protein | ||||
| STRING | EOY33703 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM877 | 26 | 107 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G58620.1 | 0.0 | C3H family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG041605t1 |
| Entrez Gene | 18590481 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



