 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG001039t3 | ||||||||
| Common Name | TCM_001039 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 318aa MW: 35828.3 Da PI: 4.6335 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.5 | 5.5e-19 | 44 | 97 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ +q+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
Thecc1EG001039t3 44 KKRRLSVDQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWK 97
566899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 128.8 | 2.3e-41 | 43 | 135 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekkrrls +qvk+LE++Fe e+kLeperKv+la+eLglqprqvavWFqnrRAR+ktkqlE+dy Lk++y++lk + ++L++++e+L +e++e
Thecc1EG001039t3 43 EKKRRLSVDQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWKTKQLERDYGLLKTSYETLKVNYDTLQHDNEALLKEIRE 135
69*************************************************************************************999986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.65E-19 | 30 | 101 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.346 | 39 | 99 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.1E-17 | 42 | 103 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 3.6E-16 | 44 | 97 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.21E-16 | 44 | 100 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 9.5E-20 | 46 | 105 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 8.9E-6 | 70 | 79 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 74 | 97 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 8.9E-6 | 79 | 95 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 4.4E-16 | 99 | 140 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008483 | Molecular Function | transaminase activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 318 aa Download sequence Send to blast |
MSICPTTDEH SPRNNHIYSR EFQSMLDGLD EEGCVEESGH VAEKKRRLSV DQVKALEKNF 60 EVENKLEPER KVKLAQELGL QPRQVAVWFQ NRRARWKTKQ LERDYGLLKT SYETLKVNYD 120 TLQHDNEALL KEIRELKAKL NGESTESNLS VKEEVIVHET DNKTLEQSEP PPVSSLVTSS 180 EPAELNYESF NNSIGSVGAT LFPDLKDGSS DSDSSAILNE DNNNCSPNNA AISSSGVLQS 240 QQHLLMSPTT TSSLNFNSSS SSPSSMNCFQ FSKSTYQPSH QYVKMEEHNF FSADEACNFF 300 SDEQAPSLHW YSPEQWN* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 91 | 99 | RRARWKTKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00051 | PBM | Transfer from AT4G40060 | Download |
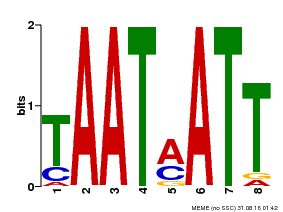
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HQ527086 | 3e-40 | HQ527086.1 Gossypium herbaceum clone NBRI_D2_391 simple sequence repeat marker, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007047858.2 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-6 | ||||
| TrEMBL | A0A061DJ94 | 0.0 | A0A061DJ94_THECC; Alanine--glyoxylate aminotransferase 2 isoform 1 | ||||
| STRING | EOX92016 | 0.0 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G40060.1 | 5e-70 | homeobox protein 16 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG001039t3 |
| Entrez Gene | 18611510 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



