 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG000235t1 | ||||||||
| Common Name | TCM_000235 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 493aa MW: 55886.8 Da PI: 5.7774 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 179.1 | 1.2e-55 | 161 | 290 | 1 | 129 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpk...kvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdk 90
+ppGfrFhPtdeelv +yL+kkv+++k++l +vi+++d+y++ePwdL++ ++e++ewyfFs++dkky+tg+r+nrat +g+Wkatg+dk
Thecc1EG000235t1 161 VPPGFRFHPTDEELVGYYLRKKVASQKIDL-DVIRDIDLYRIEPWDLQErcrIGYEEQNEWYFFSHKDKKYPTGTRTNRATMAGFWKATGRDK 252
69****************************.9***************953432233667********************************** PP
NAM 91 evlskkgelvglkktLvfykgrapkgektdWvmheyrle 129
+v++ k++l+g++ktLvfykgrap+g+ktdW+mheyrle
Thecc1EG000235t1 253 AVYD-KSKLIGMRKTLVFYKGRAPNGQKTDWIMHEYRLE 290
****.999*****************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 6.54E-61 | 156 | 310 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 59.711 | 161 | 310 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.1E-28 | 162 | 289 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0048759 | Biological Process | xylem vessel member cell differentiation | ||||
| GO:1901348 | Biological Process | positive regulation of secondary cell wall biogenesis | ||||
| GO:1990110 | Biological Process | callus formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 493 aa Download sequence Send to blast |
MTGISLKVTP AENPPSGKSR VLVGGGSPVL GNRGRLGEKR KRKPCERRAE SGAGRSDSVD 60 GRSFGGNTWP SHQPCAIERS SPPSPPSNLL PLLYSLHISL SLLSLYSLIP SLLHIFFFFF 120 FFFFSFSFSS FVFFFFIASV SNELTNSPAP TDMMESMESC VPPGFRFHPT DEELVGYYLR 180 KKVASQKIDL DVIRDIDLYR IEPWDLQERC RIGYEEQNEW YFFSHKDKKY PTGTRTNRAT 240 MAGFWKATGR DKAVYDKSKL IGMRKTLVFY KGRAPNGQKT DWIMHEYRLE TDENGPPQEE 300 GWVVCRAFKK RTNGNGQSKS IEGWDSSYFY DEPSGVSSVV DPIDYISRQP QSFLPQSFLC 360 KQETEADNLN FIHSDQFVEL PQLESPSLPL IKRPSSISLI SENTSYEEEE QNRMCNTTKK 420 VTDWRALDKF VASQLSQEDR YDGEGVPSFE ANNNSDMALL LLQSSGREEG NKLNEFLNSS 480 SDCDIGICIF DK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 1e-48 | 158 | 312 | 12 | 170 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the secondary wall NAC binding element (SNBE), 5'-(T/A)NN(C/T)(T/C/G)TNNNNNNNA(A/C)GN(A/C/T)(A/T)-3', in the promoter of target genes (By similarity). Involved in xylem formation by promoting the expression of secondary wall-associated transcription factors and of genes involved in secondary wall biosynthesis and programmed cell death, genes driven by the secondary wall NAC binding element (SNBE). Triggers thickening of secondary walls (PubMed:25148240). {ECO:0000250|UniProtKB:Q9LVA1, ECO:0000269|PubMed:25148240}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00266 | DAP | Transfer from AT2G18060 | Download |
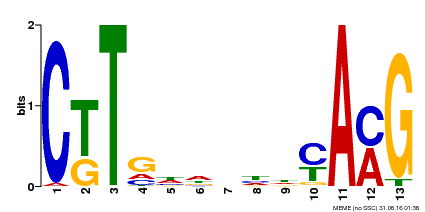
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by chitin (e.g. chitooctaose) (PubMed:17722694). Accumulates in plants exposed to callus induction medium (CIM) (PubMed:17581762). {ECO:0000269|PubMed:17581762, ECO:0000269|PubMed:17722694}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017984469.1 | 0.0 | PREDICTED: NAC domain-containing protein 37 isoform X1 | ||||
| Swissprot | Q9SL41 | 1e-165 | NAC37_ARATH; NAC domain-containing protein 37 | ||||
| TrEMBL | A0A061DFT4 | 0.0 | A0A061DFT4_THECC; Vascular related NAC-domain protein 1 isoform 1 | ||||
| STRING | EOX90892 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3686 | 26 | 61 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G18060.1 | 1e-156 | vascular related NAC-domain protein 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG000235t1 |
| Entrez Gene | 18610798 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



