 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Traes_5DL_5FD3B95C7.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 413aa MW: 45670 Da PI: 7.4372 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 29.5 | 1.3e-09 | 330 | 364 | 21 | 55 |
HSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 21 knrypsaeereeLAkklgLterqVkvWFqNrRake 55
k +yps++e+ LA+++gL+++q+ +WF N+R ++
Traes_5DL_5FD3B95C7.1 330 KWPYPSETEKIALAESTGLDQKQINNWFINQRKRH 364
569*****************************885 PP
| |||||||
| 2 | ELK | 43 | 9e-15 | 284 | 305 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELKhqLlrKY+gy gsL+qEF+
Traes_5DL_5FD3B95C7.1 284 ELKHQLLRKYGGYVGSLRQEFC 305
9********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 1.7E-20 | 151 | 195 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 5.7E-22 | 152 | 193 | IPR005540 | KNOX1 |
| SMART | SM01256 | 4.8E-28 | 201 | 252 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 5.0E-24 | 205 | 251 | IPR005541 | KNOX2 |
| PROSITE profile | PS51213 | 11.453 | 284 | 304 | IPR005539 | ELK domain |
| SMART | SM01188 | 9.7E-8 | 284 | 305 | IPR005539 | ELK domain |
| Pfam | PF03789 | 1.2E-10 | 284 | 305 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.795 | 304 | 367 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.0E-13 | 306 | 371 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 3.94E-19 | 306 | 375 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 4.07E-14 | 307 | 367 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-27 | 309 | 370 | IPR009057 | Homeodomain-like |
| Pfam | PF05920 | 2.5E-16 | 324 | 363 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 342 | 365 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 413 aa Download sequence Send to blast |
MGTKPIPSHQ PHTHSLIIHS PSGLLLLLSS SVRPPPSFVL HHFSSFQERV HLISFLVRVS 60 WPCLFLIGRF SAQIMERFPN LGGGGGSGSG SSSASMASFL QLPPPSASSP SPELAGEQHG 120 SRLALQQLLA APPPSAQQQQ QRREISPADV ATIKAKIMAH PLYSPLLASY LDCQKVGAPP 180 EVLERLSAVA AKLDAGHGRG KHESPRPDPE LDQFMEAYCN MLAKYREELA RPIQEATEFF 240 KSVETQLDSI TFTDSTNCEG AGSSEDELDT SCVEEIDPSA EDKELKHQLL RKYGGYVGSL 300 RQEFCKRRKK GKLPKEARQK LLHWWELHSK WPYPSETEKI ALAESTGLDQ KQINNWFINQ 360 RKRHWKPAPE DMPFSVMDGG VGVGVSFLPA PQGPALYMDR APFMVDGMYR LGS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during embryogenesis. {ECO:0000269|PubMed:10080693}. | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during embryogenesis. {ECO:0000269|PubMed:10080693, ECO:0000269|PubMed:9869405}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00671 | PBM | Transfer from GRMZM2G135447 | Download |
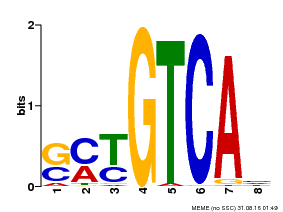
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB443455 | 0.0 | AB443455.1 Triticum aestivum WRS1 mRNA for homeobox protein, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020156617.1 | 0.0 | homeobox protein rough sheath 1-like | ||||
| Swissprot | O65034 | 1e-116 | KNOSC_ORYSI; Homeobox protein knotted-1-like 12 | ||||
| Swissprot | O80416 | 1e-116 | KNOSC_ORYSJ; Homeobox protein knotted-1-like 12 | ||||
| TrEMBL | A0A3B6MZX2 | 0.0 | A0A3B6MZX2_WHEAT; Uncharacterized protein | ||||
| STRING | Traes_5DL_5FD3B95C7.1 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4061 | 38 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 4e-95 | KNOTTED-like from Arabidopsis thaliana | ||||



