 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Traes_5BL_B2792D532.2 | ||||||||
| Common Name | WRS1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 254aa MW: 28806.7 Da PI: 5.928 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 30.3 | 7e-10 | 173 | 207 | 21 | 55 |
HSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 21 knrypsaeereeLAkklgLterqVkvWFqNrRake 55
k +yps++e+ LA+++gL+++q+ +WF N+R ++
Traes_5BL_B2792D532.2 173 KWPYPSETEKIALAESTGLDQKQINNWFINQRKRH 207
569*****************************885 PP
| |||||||
| 2 | ELK | 43.9 | 4.9e-15 | 127 | 148 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELKhqLlrKY+gy gsL+qEF+
Traes_5BL_B2792D532.2 127 ELKHQLLRKYGGYVGSLRQEFC 148
9********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03790 | 2.4E-17 | 1 | 36 | IPR005540 | KNOX1 |
| SMART | SM01255 | 1.5E-12 | 1 | 38 | IPR005540 | KNOX1 |
| SMART | SM01256 | 4.8E-28 | 44 | 95 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 2.4E-24 | 48 | 94 | IPR005541 | KNOX2 |
| PROSITE profile | PS51213 | 11.453 | 127 | 147 | IPR005539 | ELK domain |
| Pfam | PF03789 | 6.3E-11 | 127 | 148 | IPR005539 | ELK domain |
| SMART | SM01188 | 9.7E-8 | 127 | 148 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.795 | 147 | 210 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.63E-19 | 149 | 219 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 3.0E-13 | 149 | 214 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.10E-15 | 150 | 210 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 5.6E-28 | 152 | 213 | IPR009057 | Homeodomain-like |
| Pfam | PF05920 | 1.3E-16 | 167 | 206 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 185 | 208 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 254 aa Download sequence Send to blast |
MAHPLYSPLL ASYLDCQKVG APPEVLERLS AVAAKLDAGH GRGKHESPRP DPELDQFMEA 60 YCNMLAKYRE ELARPIQEAT EFFKSVETQL DSITFTDSTN CEGAGSSEDE LDTSCVEEID 120 PSAEDKELKH QLLRKYGGYV GSLRQEFCKR RKKGKLPKEA RQKLLHWWEL HSKWPYPSET 180 EKIALAESTG LDQKQINNWF INQRKRHWKP APEDMPFSVM DGGVGVSFLP APQGPALYMD 240 RAPFMVDGMY RLGS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Plays a possible role in patterning the placement of lateral organs along the axis of the shoot. Mutations in RS1 alters cell fate and causes unregulated cell division and expansion in the leaf. Probably binds to the DNA sequence 5'-TGAC-3'. | |||||
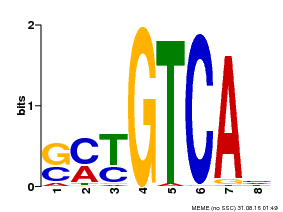
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00671 | PBM | Transfer from GRMZM2G135447 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB443455 | 0.0 | AB443455.1 Triticum aestivum WRS1 mRNA for homeobox protein, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020156617.1 | 0.0 | homeobox protein rough sheath 1-like | ||||
| Swissprot | Q41853 | 1e-111 | RSH1_MAIZE; Homeobox protein rough sheath 1 | ||||
| TrEMBL | A0A3B6LSJ7 | 0.0 | A0A3B6LSJ7_WHEAT; Uncharacterized protein | ||||
| STRING | Traes_5BL_B2792D532.2 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4061 | 38 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 3e-94 | KNOTTED-like from Arabidopsis thaliana | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



