 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | TRAES3BF062500050CFD_t1 | ||||||||
| Common Name | TRAES_3BF062500050CFD_c1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 503aa MW: 54275.5 Da PI: 6.2003 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 78 | 1.2e-24 | 204 | 259 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvk.gLtlehvkSHLQkYRla 56
k +++WtpeLH+rFv+aveqL G +kA+P++ilelm+ + Lt+++++SHLQkYR++
TRAES3BF062500050CFD_t1 204 KVKVDWTPELHRRFVQAVEQL-GLDKAVPSRILELMGNEyRLTRHNIASHLQKYRSH 259
5689*****************.**************88747**************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.428 | 201 | 261 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-24 | 202 | 262 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 7.35E-16 | 203 | 262 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.7E-24 | 204 | 259 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.9E-9 | 207 | 257 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 503 aa Download sequence Send to blast |
MLEVATLQSP NPAVFSLGEH HHVDVGFPEV TVEEDDFLLD YIDFSTCDMP FFHVDDGDDI 60 LPDLEVDPTE LLAEFADEPA TTTVLSPAAE PAPDGCETHH GGEAKTAAET ELPAEMGMEL 120 PEGKGETKGL LSSEEKDVRQ HNDNNKNNNN VGDEVCSAVT TDDSSAVVGS ENSKSSASAE 180 GHSKMTSKPA SAAAATKSSH GRRKVKVDWT PELHRRFVQA VEQLGLDKAV PSRILELMGN 240 EYRLTRHNIA SHLQKYRSHR KHLMAREAGA ARWTQKRQMY AAAGGPRKEA AAGGGPWVVP 300 TVGFPPPGTM AHAGMAHHPG QPPPFCRPLH VWGHPTGVDA PLPLSPPSML PVWPRHLAPP 360 PAWAHQPPMD PVYWHQQYNA ARKWGPQAVT QCVPPTMPPA PMMQRFAAPP MPGMMPHPMY 420 RPIPPPPSPV PQSNKVAGLQ LQLDAHPSKE SIDAAIGDVL VKPWLPLPLG LKPPSLDSVM 480 SELHKQGIPK VPPAATTNCD GAA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that promotes chloroplast development. Acts as an activator of nuclear photosynthetic genes involved in chlorophyll biosynthesis, light harvesting, and electron transport (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
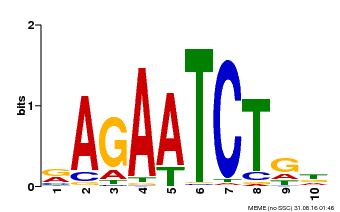
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. {ECO:0000269|PubMed:11340194}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353571 | 0.0 | AK353571.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1001B16. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020187877.1 | 0.0 | probable transcription factor GLK2 | ||||
| Swissprot | Q5NAN5 | 1e-125 | GLK2_ORYSJ; Probable transcription factor GLK2 | ||||
| TrEMBL | A0A077RRG8 | 0.0 | A0A077RRG8_WHEAT; Uncharacterized protein | ||||
| STRING | Traes_3B_73B7FB5C1.2 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2415 | 38 | 86 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 8e-56 | GBF's pro-rich region-interacting factor 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



