 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sevir.9G076700.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 379aa MW: 38495.6 Da PI: 7.9344 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.7 | 4.5e-32 | 113 | 166 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
pr+rWt+ LH+rFv+ave LGG+e+AtPk++lelm+vk+Ltl+hvkSHLQ+YR+
Sevir.9G076700.1.p 113 PRMRWTTALHARFVHAVELLGGHERATPKSVLELMNVKDLTLAHVKSHLQMYRT 166
9****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51257 | 5 | 1 | 24 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 7.2E-29 | 111 | 167 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 9.5E-16 | 111 | 167 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.3E-23 | 113 | 166 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 5.8E-8 | 114 | 165 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005618 | Cellular Component | cell wall | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 379 aa Download sequence Send to blast |
MLQRNLPNLS LGISPPAASS AAACDALAVA RPPLAEPSTA PHAEGSGEVG FFANPSPVAE 60 PPGLSLGLGT PAARGGDAAG RRGHLEPQGC AFKRAATRAS LPGGSTKRSA RAPRMRWTTA 120 LHARFVHAVE LLGGHERATP KSVLELMNVK DLTLAHVKSH LQMYRTVKST DRSLHIATGE 180 ALPLQRTTAT GMEAAAAAAA AGGGAAAGGG VVVVPVPAAC DDMVGICSSP SAGSAPPAAA 240 TTSAAHFLYA PPATAPLAAA PPPPPPIPPR TDHAPGVLGK GAAIVDSAHR CQKHNFSPPA 300 LQDTQAAQEE VNRHLAMGLH ARSEAVATNC SSPASSSSPS LASVELLADD MYAPNLEISL 360 GRQDWSMERP EELSLKYL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 3e-16 | 114 | 168 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 3e-16 | 114 | 168 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 2e-16 | 114 | 168 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-16 | 114 | 168 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-16 | 114 | 168 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-16 | 114 | 168 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 3e-16 | 114 | 168 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 3e-16 | 114 | 168 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 3e-16 | 114 | 168 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 3e-16 | 114 | 168 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 3e-16 | 114 | 168 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 3e-16 | 114 | 168 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00107 | PBM | Transfer from AT5G42630 | Download |
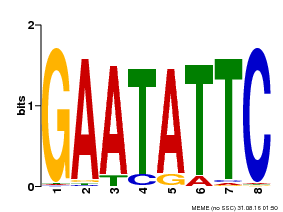
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sevir.9G076700.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT065124 | 1e-151 | BT065124.1 Zea mays full-length cDNA clone ZM_BFb0205B22 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004981643.1 | 0.0 | probable transcription factor RL9 | ||||
| TrEMBL | K4ABD0 | 0.0 | K4ABD0_SETIT; Uncharacterized protein | ||||
| STRING | Si036187m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6554 | 37 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42630.2 | 5e-37 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sevir.9G076700.1.p |



