 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sevir.4G282500.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 433aa MW: 46842.2 Da PI: 9.4811 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130.2 | 7.4e-41 | 160 | 236 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
CqvegC++dls+ak yhr+hkvCe h+kap+v+v+gle+rfCqqCsrfh l efD++krsCrrrL +hn+rrrk+q+
Sevir.4G282500.2.p 160 CQVEGCKVDLSSAKAYHRKHKVCEDHAKAPKVVVAGLERRFCQQCSRFHGLAEFDQNKRSCRRRLTHHNARRRKPQT 236
**************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.2E-33 | 153 | 221 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.146 | 157 | 234 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.09E-38 | 159 | 240 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 5.3E-32 | 160 | 233 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 433 aa Download sequence Send to blast |
MGSFGMNWDQ KNSMVWDCEN LAPSVPNEIV RHGSANSSGG TLTSSSELGH GSSKSSISGS 60 IDSPFGVGNS IEFNFAAVER HVKDMGKNGR VDDSRTSPSS MIAFSHGEPS ISLKLGKRAY 120 VENVCGRQDN KSSAPSTVTS ASTVVKKTKV SHQNVKNSYC QVEGCKVDLS SAKAYHRKHK 180 VCEDHAKAPK VVVAGLERRF CQQCSRFHGL AEFDQNKRSC RRRLTHHNAR RRKPQTDTIS 240 FNSSRLSTMF YDTSQQTNLF FSQPLFSQVR SNALSSWDNL GGFKFVETKH MSMHPMKTVG 300 LDELPFSNLQ ISTSVAAQTA RHHNFDGLMP VKGTNTKVLN QGVEASTAAS NSNGAPELGR 360 ALSLLSDGSW GSSSTVIQQH NSHVHTGAMP PLGTIAVSNP VTNHLDPSPG GFWHDDPATL 420 DGTLQIQAST HL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-28 | 160 | 233 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00161 | DAP | Transfer from AT1G27360 | Download |
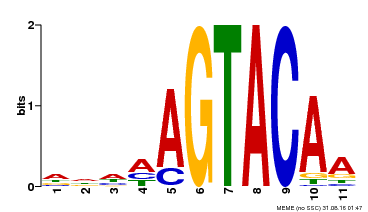
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sevir.4G282500.2.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156b and miR156h. {ECO:0000305|PubMed:16861571}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004966343.1 | 0.0 | squamosa promoter-binding-like protein 12 isoform X1 | ||||
| Refseq | XP_004966344.1 | 0.0 | squamosa promoter-binding-like protein 12 isoform X1 | ||||
| Refseq | XP_004966345.1 | 0.0 | squamosa promoter-binding-like protein 12 isoform X1 | ||||
| Refseq | XP_022681748.1 | 0.0 | squamosa promoter-binding-like protein 12 isoform X1 | ||||
| Swissprot | A2YGR5 | 1e-165 | SPL12_ORYSI; Squamosa promoter-binding-like protein 12 | ||||
| TrEMBL | K3XX13 | 0.0 | K3XX13_SETIT; Uncharacterized protein | ||||
| STRING | Si006471m | 0.0 | (Setaria italica) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G27360.4 | 3e-37 | squamosa promoter-like 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sevir.4G282500.2.p |



