 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PGSC0003DMP400029334 | ||||||||
| Common Name | LOC102592561 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 326aa MW: 36691 Da PI: 4.3953 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.8 | 4.5e-19 | 62 | 115 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k++++ +q+++Le+ Fe +++++ e++ +LA++lgL+ rqV +WFqNrRa++k
PGSC0003DMP400029334 62 KKRRLRVDQVKALEKNFEVDNKLDPERKMKLAQELGLQPRQVAIWFQNRRARWK 115
45578899*********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 126.5 | 1.1e-40 | 61 | 153 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLre 89
ekkrrl +qvk+LE++Fe ++kL+perK++la+eLglqprqva+WFqnrRAR+ktkqlE+dy++Lk+++d+lk++ e++++++e+L +
PGSC0003DMP400029334 61 EKKRRLRVDQVKALEKNFEVDNKLDPERKMKLAQELGLQPRQVAIWFQNRRARWKTKQLERDYNVLKANFDSLKHNYESIQHDNEALLK 149
69*************************************************************************************99 PP
HD-ZIP_I/II 90 elke 93
e++e
PGSC0003DMP400029334 150 EINE 153
9976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 17.313 | 57 | 117 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 2.35E-19 | 59 | 119 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 3.5E-18 | 60 | 121 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 2.6E-16 | 62 | 115 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.31E-16 | 62 | 118 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.2E-20 | 69 | 116 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 8.3E-6 | 88 | 97 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 92 | 115 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 8.3E-6 | 97 | 113 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 8.6E-18 | 117 | 159 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 326 aa Download sequence Send to blast |
MKRVRSNDSL GAPLTSICQN TTDEHNNWNN HVYSRDFQSM LEGLDDDEAT TCVEESGVGS 60 EKKRRLRVDQ VKALEKNFEV DNKLDPERKM KLAQELGLQP RQVAIWFQNR RARWKTKQLE 120 RDYNVLKANF DSLKHNYESI QHDNEALLKE INELKSKLQG GNNEIKIAVK EEAFESESDD 180 KETEQSNQCN NSNTELNFDT IIMNPSIIFT DFKDGSSDSD SSAILNEDNA VISSSEAFLI 240 KSGSGSGSGS GSGSSSPDSN SSLNCFQFLE SNPKSILTNE VDFSSSQFVK LEEHNFFSGE 300 ESCSTLLFTD EQAPTLHWYC SEDWN* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 109 | 117 | RRARWKTKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
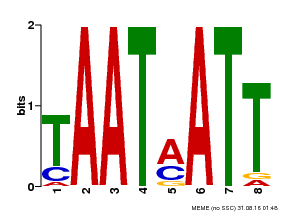
| Motif ID | Method | Source | Motif file |
| MP00051 | PBM | Transfer from AT4G40060 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PGSC0003DMP400029334 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975444 | 1e-151 | HG975444.1 Solanum pennellii chromosome ch05, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006366219.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-6-like | ||||
| TrEMBL | M1BEC0 | 0.0 | M1BEC0_SOLTU; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400043251 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2718 | 24 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G40060.1 | 9e-68 | homeobox protein 16 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | PGSC0003DMP400029334 |
| Entrez Gene | 102592561 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



