 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PGSC0003DMP400028208 | ||||||||
| Common Name | LOC102586298 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 349aa MW: 39105.7 Da PI: 4.3154 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.3 | 1.2e-18 | 57 | 110 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
PGSC0003DMP400028208 57 KKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWK 110
456899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 130.5 | 6.7e-42 | 56 | 148 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLre 89
+kkrrls eqvk+LE++Fe e+kLeperKv+la+eLglqprqvavWFqnrRAR+ktkqlE+dy++Lk+++d+lk++ e+L++++e+L +
PGSC0003DMP400028208 56 QKKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWKTKQLERDYNVLKANFDSLKHNYESLKHDNEALGK 144
69*************************************************************************************99 PP
HD-ZIP_I/II 90 elke 93
e++e
PGSC0003DMP400028208 145 EIRE 148
9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 17.88 | 52 | 112 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 3.29E-19 | 53 | 114 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 4.6E-18 | 55 | 116 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.08E-17 | 56 | 113 | No hit | No description |
| Pfam | PF00046 | 8.1E-16 | 57 | 110 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 8.5E-20 | 59 | 119 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.0E-5 | 83 | 92 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 87 | 110 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.0E-5 | 92 | 108 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 9.4E-17 | 112 | 152 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 349 aa Download sequence Send to blast |
MKRLSSSDSL GALMSMCPST TDDQNHGNSH VYSTRDFQSM LELGLDEEGC VEESGQKKRR 60 LSVEQVKALE KNFEVENKLE PERKVKLAQE LGLQPRQVAV WFQNRRARWK TKQLERDYNV 120 LKANFDSLKH NYESLKHDNE ALGKEIRELK LKVYGGENGE SRGAVVVSVK EEAMESENDD 180 KMIEQNKPND LLEEEEDEDQ DQDDDVEINA TIASTIFADF NKDGSSDSDN SSAILNEDNS 240 PNAAAISSSG AFLISNDGGV GCSSPLNFTF KFTESSPKSI LGDSQKANCF TYQPSSTTTT 300 TQYVKMEEHN FFNGEESCST LFSDEQAPTL QWYCSEDWNF KDTPSHLS* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 104 | 112 | RRARWKTKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00271 | DAP | Transfer from AT2G22430 | Download |
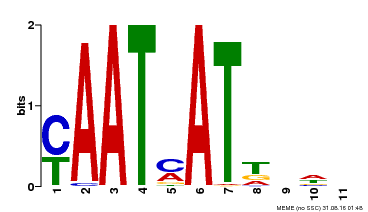
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PGSC0003DMP400028208 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975523 | 0.0 | HG975523.1 Solanum lycopersicum chromosome ch11, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006353200.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-6 | ||||
| TrEMBL | M1BBQ5 | 0.0 | M1BBQ5_SOLTU; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400041655 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2718 | 24 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22430.1 | 5e-63 | homeobox protein 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | PGSC0003DMP400028208 |
| Entrez Gene | 102586298 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



