 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PGSC0003DMP400026610 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 655aa MW: 73804.3 Da PI: 9.7269 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.3 | 0.00051 | 538 | 563 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y C C++sFs+k +L+ H ++ +
PGSC0003DMP400026610 538 YHCDleGCSMSFSSKQELTLHKKNvC 563
889999***************99866 PP
| |||||||
| 2 | zf-C2H2 | 12 | 0.00065 | 562 | 585 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirtH 23
+Cp C+k F ++ +L++H r+H
PGSC0003DMP400026610 562 VCPveGCKKKFFSHKYLVQHRRVH 585
69999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 11.9 | 0.00071 | 621 | 647 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y C+ Cg++F+ s++ rH r+ H
PGSC0003DMP400026610 621 YACTetGCGQTFRFVSDFSRHKRKtgH 647
889999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 8.6 | 538 | 560 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.012 | 561 | 585 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.572 | 561 | 590 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.0E-5 | 561 | 584 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 563 | 585 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 6.35E-10 | 577 | 619 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.2E-5 | 585 | 597 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0014 | 591 | 615 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.741 | 591 | 620 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 593 | 615 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 6.1E-4 | 598 | 613 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 3.78E-8 | 609 | 643 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.9E-9 | 614 | 644 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.36 | 621 | 647 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.157 | 621 | 652 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 623 | 647 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 655 aa Download sequence Send to blast |
MVKLYVLIAG RFVKRAPALI DVPVESSDRI QKLNNGSVEG FSRTKAHKET SSLGLLALAY 60 ANSSDSDEDE VEADIPVEAC ESRHTDSEDE VFLRVIDPYG NHRQKRAVSQ GRNCQKTDNS 120 VQLENESYPS GESNTLLGRS SHQPRSHQVA AKCISNIGEI VQNNAVAPFD HARMQFTSTS 180 DEDSFRIHVF CLQHAVQVEE QLRRIGGARI SLLCHPDYPK LEAQAKQVAE ELGSDHFWRE 240 ISFREATKDD EEMIQSALEI EEAIHGNGDW TVKLDINLFY SANLSRSPLY SKQMPYNFII 300 YNAFGRNSPD NTPEKSEYTG RGSGKQRRAI VAGKWCGKVW MSSQVHPLLA ERTIDEEQEQ 360 NKSISAQIKI EVKSERPRER TPTGKTVSTA CKTGKKRSST AVSRNASNAQ LIIADDHDDS 420 LLSSILQQHR KTNLRSKRIK YETPEPQKDV DKKKIFGSII DDDPDGGPST RLRKRIPKPS 480 NESPAKLVKV KPAPTKQHES KKGPKVKLPS ANSNAKKEPV TKGPRSNIGK RMREEGEYHC 540 DLEGCSMSFS SKQELTLHKK NVCPVEGCKK KFFSHKYLVQ HRRVHMDDRP LKCPWKGCKM 600 TFKWAWARTE HIRVHTGARP YACTETGCGQ TFRFVSDFSR HKRKTGHVSK KGRS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 6e-65 | 538 | 651 | 23 | 136 | Lysine-specific demethylase REF6 |
| 6a58_A | 6e-65 | 538 | 651 | 23 | 136 | Lysine-specific demethylase REF6 |
| 6a59_A | 6e-65 | 538 | 651 | 23 | 136 | Lysine-specific demethylase REF6 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
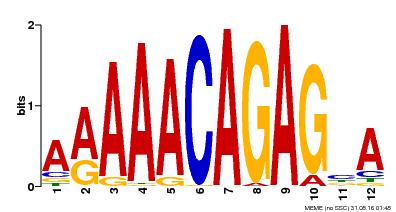
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PGSC0003DMP400026610 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975442 | 0.0 | HG975442.1 Solanum pennellii chromosome ch03, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006351452.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 | ||||
| TrEMBL | M1B7Y4 | 0.0 | M1B7Y4_SOLTU; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400039224 | 0.0 | (Solanum tuberosum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 9e-69 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | PGSC0003DMP400026610 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



