 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PGSC0003DMP400002220 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 283aa MW: 33009.1 Da PI: 7.263 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 158.2 | 3.5e-49 | 19 | 144 | 2 | 128 |
NAM 2 ppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdk 90
pGfrFhPtdeelv +yL++kve+k++++ e ik++diyk++PwdLpk v e+ke+y+F+ r +ky+++ r+nr+tksg+Wkatg dk
PGSC0003DMP400002220 19 LPGFRFHPTDEELVGFYLRRKVENKRINM-ELIKHIDIYKYDPWDLPKLV--EDKEMYYFCMRGRKYKNSVRPNRVTKSGFWKATGIDK 104
69***************************.99***************655..789********************************** PP
NAM 91 evlskkgel..vglkktLvfykgrapkgektdWvmheyrl 128
+v+s++++ +glkk Lv+y+g+a kg+ktdW+mhe+rl
PGSC0003DMP400002220 105 PVYSSTTQCiiIGLKKCLVYYRGSAGKGTKTDWMMHEFRL 144
****74443559**************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 7.45E-54 | 13 | 165 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 51.807 | 18 | 165 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.4E-24 | 20 | 144 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005992 | Biological Process | trehalose biosynthetic process | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006561 | Biological Process | proline biosynthetic process | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0010120 | Biological Process | camalexin biosynthetic process | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 283 aa Download sequence Send to blast |
MADEIKTMSK EGNEEDVILP GFRFHPTDEE LVGFYLRRKV ENKRINMELI KHIDIYKYDP 60 WDLPKLVEDK EMYYFCMRGR KYKNSVRPNR VTKSGFWKAT GIDKPVYSST TQCIIIGLKK 120 CLVYYRGSAG KGTKTDWMMH EFRLPPTSNT NHQAEVWTLC RIFKRSTSTY KRSSYSTQEV 180 KHLLGDASSK ACSMESDENS TDHDHSNSFK DISFQQQNYN NYEAQMDEIR NQQNLISHDS 240 TFPSSSSISF WNTYSTQEEY LFGDHGNKWD DLKSIVDLAT MM* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 1e-47 | 18 | 171 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 1e-47 | 18 | 171 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 1e-47 | 18 | 171 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 1e-47 | 18 | 171 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 1e-47 | 18 | 171 | 17 | 171 | NAC domain-containing protein 19 |
| 4dul_B | 1e-47 | 18 | 171 | 17 | 171 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
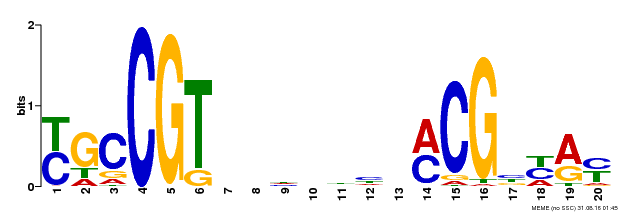
| UniProt | Transcription factor that binds to the 5'- RRYGCCGT-3' consensus core sequence. Central longevity regulator. Negative regulator of leaf senescence. Modulates cellular H(2)O(2) levels and enhances tolerance to various abiotic stresses through the regulation of DREB2A. {ECO:0000269|PubMed:22345491}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00310 | DAP | Transfer from AT2G43000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | PGSC0003DMP400002220 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by H(2)O(2), paraquat, ozone, 3-aminotriazole and salt stress. {ECO:0000269|PubMed:22345491}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HG975444 | 1e-168 | HG975444.1 Solanum pennellii chromosome ch05, complete genome. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015159594.1 | 0.0 | PREDICTED: transcription factor JUNGBRUNNEN 1-like | ||||
| Swissprot | Q9SK55 | 2e-89 | NAC42_ARATH; Transcription factor JUNGBRUNNEN 1 | ||||
| TrEMBL | M0ZL44 | 0.0 | M0ZL44_SOLTU; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400003082 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1697 | 24 | 69 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43000.1 | 4e-86 | NAC domain containing protein 42 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | PGSC0003DMP400002220 |



