 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0486s0080.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1691aa MW: 190630 Da PI: 8.9982 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.7 | 0.00038 | 1599 | 1621 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
SapurV1A.0486s0080.1.p 1599 CPvkGCGKKFFSHKYLVQHRRVH 1621
9999*****************99 PP
| |||||||
| 2 | zf-C2H2 | 11.6 | 0.00088 | 1657 | 1683 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
SapurV1A.0486s0080.1.p 1657 YVCAeeGCGQTFRFVSDFSRHKRKtgH 1683
89********************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 4.8E-14 | 29 | 68 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.495 | 30 | 71 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 3.9E-14 | 31 | 64 | IPR003349 | JmjN domain |
| SuperFamily | SSF51197 | 6.32E-27 | 118 | 177 | No hit | No description |
| SMART | SM00558 | 1.2E-50 | 196 | 365 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 34.251 | 196 | 365 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 6.32E-27 | 214 | 379 | No hit | No description |
| Pfam | PF02373 | 9.5E-37 | 229 | 348 | IPR003347 | JmjC domain |
| SMART | SM00355 | 13 | 1574 | 1596 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.0045 | 1597 | 1621 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.383 | 1597 | 1626 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.2E-5 | 1598 | 1620 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1599 | 1621 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.02E-9 | 1613 | 1655 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.5E-8 | 1621 | 1648 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0014 | 1627 | 1651 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.741 | 1627 | 1656 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1629 | 1651 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 6.48E-8 | 1645 | 1679 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.7E-9 | 1649 | 1680 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.85 | 1657 | 1683 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.24 | 1657 | 1688 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1659 | 1683 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1691 aa Download sequence Send to blast |
MAAPSGLSLD PPQPPTTTEV CQWLKNLPLA PEYRPTLSEF QDPIAYIFKI EKEASQYGIC 60 KIIPPVLPSA KKTTLSNLNR SLCARNGGSS APTFTTRQQQ IGFCPRKPRP VQKPVWQSGE 120 TYTFQEFETK ARTFEKNYLK KFSKKGALSP LEFETLYWKA TLDKPFSVEY ANDMPGSAFS 180 PRKKEGQGGV AGDGMSVGET EWNMRGVSRA KGSLLRFMKE EIPGVTSPMV YIGMMFSWFA 240 WHVEDHDLHS LNYMHMGAGK TWYGVPREAA VAFEEVVRVH GYGGEINPLV TFSVLGEKTT 300 VMSPEVFISA GVPCCRLVQN AGEFVVTFPR AYHSGFSHGF NCGEAANIAT PEWLMVAKDA 360 AIRRASINYP PMVSHFQLLY DLALEFCTRI PVNISAKPRS SRLKDKQKGE GEMLVKEQFV 420 KNMIQNNDLL HSLGKGSSVV LLPRSSSDIS VCSKLRVGSQ LRDSPTLGLC SQKDVMKSSK 480 SSGSGDILQD KDQEINQVKG FFSVKAKFAS LCERNRFSIL NGNECSERMN IGTGRGSSIH 540 GDKLSDQRLF SCVTCGVLSF DCLAIIQPKE AASRYLMSAD CSFFNDWAIG SGVTRDVFAV 600 AGGIANISEQ NSSRWVEKNT TAGFYDVPVQ SPNYHIQMAD QGVEVASSSA KQLEASALGL 660 LALNYGNSSD SEEDQVEADL SHHDEIKMTN YSLENKYQCQ SSASPTSKQK YCDAATGGLP 720 QSPSSLDEQD DVPLKANDMY PEHGDRGDNL KDKTDDTLKS SFGFPTGNPA SIESNILDGR 780 YTDPMSMPHV SLNCSPIVHD AERTKFNRPI APLEKPDMPF TQRSDKESSC MHVFCLEHAV 840 EIEQQLSQIG GVHILLLCHP EYPRIEREAK LVSEELGIDY LWNDITFRDA AKKDEEWIQS 900 ALDSEVAIPG NGDWAVKLGI NLFYSANLSR SPFYSKQMPY NSVIYNAFGL ASSVSSPPKF 960 KVYGRRSGKP KKVVAGKWCG KVWMSNQVHP FLAIRDHMDQ DHEQEQERSF HALATPDEKL 1020 EKKPQTSNRT ETARKSGKKR KITAGSRSIK KVKCLEAEEP DSEDSMGDNC HGQHVRVHNS 1080 KNNEDAEREI SYDLVPDSHQ QHGRCRRKWA KSVESDDAVS DDPLAEHVRQ QYRRMRRSKQ 1140 DKPIKRENTA SYASVENKFQ KQLKRVHRSD RAKFFKRQSV ASDDSLDGNS DQWHERATRR 1200 TQAKYAESED AISDDSPEES SRWLHGRVPK SKQLKYIDEE GAISDDSLES GSHQHNRRVS 1260 RGTRVQLIKR NDMVSDDSLD ESSYQRLPRF SRSKLAKLIE RENAVSDDSL DGNIHQQHGR 1320 ILKSKQAKFV EGEDAISDDS LEDNTDWQRK RIPRSKMAKF VESEDAASDD LQEEDTHQHR 1380 RRIPKSKRAN FIEREVAISD DLWGNNAHRH SRKTPRSKQA RFIEREDLVS DDLLEDDSDQ 1440 QEKRILRSKQ KKSATFCQMK RGIPRKPKCI APKMMKKETL QSMKQERRIK QETPQLRIGK 1500 SELNARQFDL RAEEGLEGGP STRLRKRPSK PPKQSGTKLK EKQQNSRKKL KDASAAKAPV 1560 GRKNVKIKDE EAEYQCDIDG CTMGFGSKQE LAMHKRNICP VKGCGKKFFS HKYLVQHRRV 1620 HIDDRPLKCP WKGCKMTFKW AWARTEHIRV HTGARPYVCA EEGCGQTFRF VSDFSRHKRK 1680 TGHSAKKGRG * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 3e-79 | 18 | 493 | 3 | 457 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 3e-79 | 18 | 493 | 3 | 457 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1524 | 1550 | RKRPSKPPKQSGTKLKEKQQNSRKKLK |
| 2 | 1537 | 1548 | KLKEKQQNSRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
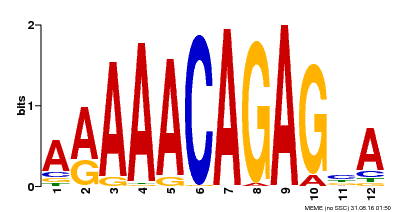
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0486s0080.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011024100.1 | 0.0 | PREDICTED: lysine-specific demethylase JMJ705 isoform X1 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A2K1XK92 | 0.0 | A0A2K1XK92_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0015s10040.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5648 | 32 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0486s0080.1.p |



