 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0403s0170.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1093aa MW: 123386 Da PI: 5.7468 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 178 | 1.2e-55 | 20 | 136 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahs 88
++ ++rwl++ ei++iL n++k++++ e+++ p+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+++vl+cyYah+
SapurV1A.0403s0170.1.p 20 VEAQNRWLRPAEIVEILSNYQKFRIAPEPAHMPSSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKCGSIDVLHCYYAHG 106
6679*********************************************************************************** PP
CG-1 89 eenptfqrrcywlLeeelekivlvhylevk 118
e+n++fqrr+yw+Leeel++ivlvhy+evk
SapurV1A.0403s0170.1.p 107 EDNENFQRRSYWMLEEELSHIVLVHYREVK 136
***************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 80.494 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 5.5E-76 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.8E-48 | 21 | 134 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF01833 | 5.9E-5 | 513 | 596 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 6.02E-15 | 513 | 599 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:1.25.40.20 | 2.2E-13 | 695 | 806 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 14.663 | 696 | 815 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 8.27E-10 | 696 | 803 | No hit | No description |
| SuperFamily | SSF48403 | 2.21E-12 | 702 | 805 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF52540 | 1.14E-7 | 914 | 968 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 2 | 917 | 939 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0046 | 920 | 938 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.206 | 920 | 947 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0039 | 940 | 962 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.597 | 941 | 965 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.2E-4 | 943 | 962 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1093 aa Download sequence Send to blast |
MADSRRYPLG NQLDIQQILV EAQNRWLRPA EIVEILSNYQ KFRIAPEPAH MPSSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHER LKCGSIDVLH CYYAHGEDNE NFQRRSYWML 120 EEELSHIVLV HYREVKGTRT NFSRVKEYEE PIPYSQETED TMPNSEMDTS VSSSFHSNGY 180 QVPTRTTDST SMNSALASEY EDAESGYNNQ ASSRFNSFFK AQKHPMEKID TGASIPYDHM 240 LFSSDYQGKL PAVSVNEFIS PAQVDTAKDI NGTQPASEPH KVLDLPSWED VLENCSRGTE 300 CMPYQTTLLS QVDTVGFIPK QEEDILEKFL TNSFNKKQDI GSHLLDQEAW QTIEGGYLHR 360 SKWSMDQKSH LDSGYDLTAR FHDQQLDSGN LINTFESFYA QENDGHIQND LQIQPANSDH 420 GMTLEGKAIH SASMKQHILE DSGTEGLKKL DSFTRWMSKE LEDVDQPHLQ SSSGTCWTMA 480 ESENVVDADN PSRGHLFTDM LGPSLSQDQL FSIIDFLPNW AYAGTEIKVL VMGRFLKCQE 540 EAENFKWSIM FGEVEVPAEF IADGVLCCTT PSHKDGRVPF YVTCSNRVAC SEVREFEYRP 600 IHLQDITNNY INSATEDLHM GLAKMLSLNS ASPSKYDSSN IEEISQLSNK ISSLLKEGNE 660 TWDQMIKLTS QEGFYSEKLK DLLLQKVLKE QLHAWLLQKV AEGGKGPSVL DEGGQGVLHF 720 AAALGYEWAL EPTVVAGVSV NFRNVNGWTA LHWAASYERT VASLIHLGAA PGALTDPTPK 780 YPTGRTPADL ASANGHKGIS GFLAESALSA HLSSLNLEKQ DGDAAESSRI PALLTVSDCN 840 ETPFNDADLP CGLSLKDSLA AVCNATQAAA RIHQVFRVQS FQKKQLKEYG DDKHGMSHER 900 ALSLIAVKSQ KAGQYDEPVH AAIRIQNKFR GWKGRKEFLI IRQRIVKIQA HVRGHQVRKN 960 YRKIIWSVGI LDKIILRWRR KGSGLRGFKP EALTEGSSMQ IVSSKDDDDD FLKEGRKQTE 1020 ERSRIALARV KSMHQHPEAR EQYHRLRNVI AEIQETKAMY EWADNSEVMA EFDDLIDLET 1080 LWDGDTFMPT DS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
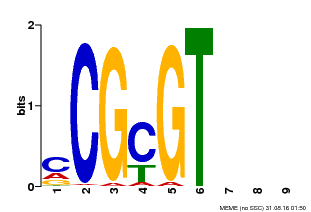
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0403s0170.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024456348.1 | 0.0 | calmodulin-binding transcription activator 3 isoform X1 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A2K2AD13 | 0.0 | A0A2K2AD13_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0007s05410.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8284 | 27 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0403s0170.1.p |



