 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | SapurV1A.0066s0010.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Salix
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 580aa MW: 64832.3 Da PI: 6.36 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 92.1 | 5.6e-29 | 60 | 144 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW++qe+ aL+++r++m+ +r ++lk+plWee+s+k++e g++rs+k+Ckek+en+ k++k++keg+ ++++++s +++fd+lea
SapurV1A.0066s0010.1.p 60 RWPRQETSALLKIRSDMDAVFRVSSLKGPLWEEISRKLAELGYHRSAKKCKEKFENVYKYHKRTKEGRTGKSEGKS--YKFFDELEA 144
8********************************************************************9877776..******985 PP
| |||||||
| 2 | trihelix | 102.2 | 4.1e-32 | 399 | 484 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+k ev+aLi++r +++ +++++ +k+plWe++s+ m++ g++r++k+Ckekwen+nk++kk+k+++kkr +e+s+tcpyfdql+a
SapurV1A.0066s0010.1.p 399 RWPKVEVQALINLRANLDIKYQENGAKGPLWEDISAGMKKIGYNRNAKRCKEKWENINKYFKKVKDSNKKR-PEDSKTCPYFDQLDA 484
8*********************************************************************8.99***********85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50090 | 7.375 | 53 | 117 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.013 | 57 | 119 | IPR001005 | SANT/Myb domain |
| CDD | cd12203 | 9.79E-24 | 59 | 124 | No hit | No description |
| Pfam | PF13837 | 5.0E-20 | 59 | 144 | No hit | No description |
| SMART | SM00717 | 0.0083 | 396 | 458 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.6E-4 | 398 | 455 | IPR009057 | Homeodomain-like |
| CDD | cd12203 | 2.42E-25 | 398 | 463 | No hit | No description |
| PROSITE profile | PS50090 | 6.946 | 398 | 456 | IPR017877 | Myb-like domain |
| Pfam | PF13837 | 8.0E-22 | 399 | 485 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 580 aa Download sequence Send to blast |
MIGDSSVQAT SSDAAATEAR VVTEGGEGGG GGAFGSNSGE EDKNMGVDHE VSRMNYGANR 60 WPRQETSALL KIRSDMDAVF RVSSLKGPLW EEISRKLAEL GYHRSAKKCK EKFENVYKYH 120 KRTKEGRTGK SEGKSYKFFD ELEAFQSHPS HSTQPTQPPP KAQTASATIT TLPWTNNTAI 180 VSHGTVPSRT NPMDIMSPSI STPTNNHTIS PMPISSNPMN PSQNAYPSLQ NLTTNLLASS 240 SPSSTASDEV LEVSKKRKRE SNWKDFFKRL TRDVIKKQED LQEKFLETME KCERERMARE 300 EAWRMQEMAR INREHETLIQ ERSTAAAKDA AVVAFLQKIS GQQDSIQTQE IPQPTTAPAP 360 PPPQPLQLRA PPTSLAPVTK LEVPKRDNGD NFTVSSPSRW PKVEVQALIN LRANLDIKYQ 420 ENGAKGPLWE DISAGMKKIG YNRNAKRCKE KWENINKYFK KVKDSNKKRP EDSKTCPYFD 480 QLDALYKEKN KMEITVSSGN AVKPPSTMEP LIVRPEQQWY FQQAARPQTI IEDSERNINI 540 DYNIEDDDGD DGDTDEEEDE GGGFEVVANK SAPLVNGDE* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds specific DNA sequence. {ECO:0000250}. | |||||
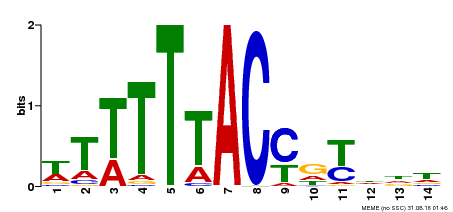
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00244 | DAP | Transfer from AT1G76890 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | SapurV1A.0066s0010.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JN315782 | 0.0 | JN315782.1 Populus tomentosa SANT DNA-binding domain-containing protein (GT2) gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002300920.2 | 0.0 | trihelix transcription factor GT-2 | ||||
| Swissprot | Q39117 | 1e-148 | TGT2_ARATH; Trihelix transcription factor GT-2 | ||||
| TrEMBL | B9GTJ1 | 0.0 | B9GTJ1_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0002s06900.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF373 | 34 | 181 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G76890.2 | 1e-115 | Trihelix family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | SapurV1A.0066s0010.1.p |



