 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT1G76890.2 | ||||||||
| Common Name | AT-GT2, F7O12.6, GT2, GT-2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 575aa MW: 65837.2 Da PI: 6.7687 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 95.5 | 5e-30 | 41 | 125 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW++ e+laL+++r+em++++r++ lk+plWee+s+km+e g++rs+k+Ckek+en+ k++k++keg+ ++++++ t+++f++lea
AT1G76890.2 41 RWPRPETLALLRIRSEMDKAFRDSTLKAPLWEEISRKMMELGYKRSSKKCKEKFENVYKYHKRTKEGRTGKSEGK--TYRFFEELEA 125
8********************************************************************975544..6******985 PP
| |||||||
| 2 | trihelix | 108.8 | 3.5e-34 | 397 | 482 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+k ev aLi++r+++e +++++ k+plWee+s+ mr+ g++rs+k+Ckekwen+nk++kk+ke++kkr + +s+tcpyf+qlea
AT1G76890.2 397 RWPKTEVEALIRIRKNLEANYQENGTKGPLWEEISAGMRRLGYNRSAKRCKEKWENINKYFKKVKESNKKR-PLDSKTCPYFHQLEA 482
8*********************************************************************8.9************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00717 | 0.086 | 38 | 100 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS50090 | 7.073 | 40 | 98 | IPR017877 | Myb-like domain |
| Pfam | PF13837 | 5.1E-20 | 40 | 125 | No hit | No description |
| CDD | cd12203 | 9.56E-25 | 40 | 105 | No hit | No description |
| PROSITE profile | PS50090 | 7.526 | 390 | 454 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 9.2E-4 | 394 | 456 | IPR001005 | SANT/Myb domain |
| Pfam | PF13837 | 7.1E-24 | 396 | 483 | No hit | No description |
| CDD | cd12203 | 3.02E-26 | 396 | 461 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 2.7E-4 | 396 | 453 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 575 aa Download sequence Send to blast |
MSGNSEGLLE SSGGGVGGSV EEEKDMKMEE TGEGAGSGGN RWPRPETLAL LRIRSEMDKA 60 FRDSTLKAPL WEEISRKMME LGYKRSSKKC KEKFENVYKY HKRTKEGRTG KSEGKTYRFF 120 EELEAFETLS SYQPEPESQP AKSSAVITNA PATSSLIPWI SSSNPSTEKS SSPLKHHHQV 180 SVQPITTNPT FLAKQPSSTT PFPFYSSNNT TTVSQPPISN DLMNNVSSLN LFSSSTSSST 240 ASDEEEDHHQ VKSSRKKRKY WKGLFTKLTK ELMEKQEKMQ KRFLETLEYR EKERISREEA 300 WRVQEIGRIN REHETLIHER SNAAAKDAAI ISFLHKISGG QPQQPQQHNH KPSQRKQYQS 360 DHSITFESKE PRAVLLDTTI KMGNYDNNHS VSPSSSRWPK TEVEALIRIR KNLEANYQEN 420 GTKGPLWEEI SAGMRRLGYN RSAKRCKEKW ENINKYFKKV KESNKKRPLD SKTCPYFHQL 480 EALYNERNKS GAMPLPLPLM VTPQRQLLLS QETQTEFETD QREKVGDKED EEEGESEEDE 540 YDEEEEGEGD NETSEFEIVL NKTSSPMDIN NNLFT |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 83 | 91 | KRSSKKCKE |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.25099 | 0.0 | root| silique | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 256332_at | 0.0 | ||||
| Expression Atlas | AT1G76890 | - | ||||
| AtGenExpress | AT1G76890 | - | ||||
| ATTED-II | AT1G76890 | - | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | encodes a plant trihelix DNA-binding protein | |||||
| UniProt | Probable transcription factor that binds specific DNA sequence. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00244 | DAP | 27203113 | Download |
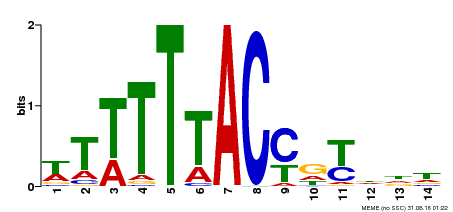
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT1G76890.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT1G76890 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT030363 | 0.0 | BT030363.1 Arabidopsis thaliana At1g76890 mRNA, complete cds. | |||
| GenBank | X72780 | 0.0 | X72780.1 A.thaliana mRNA for GT-2 factor. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_177815.1 | 0.0 | Duplicated homeodomain-like superfamily protein | ||||
| Swissprot | Q39117 | 0.0 | TGT2_ARATH; Trihelix transcription factor GT-2 | ||||
| TrEMBL | A0A178W7Z2 | 0.0 | A0A178W7Z2_ARATH; GT2 | ||||
| STRING | AT1G76890.2 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5995 | 28 | 47 | Representative plant | OGRP663 | 15 | 73 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT1G76890.2 |
| Entrez Gene | 844024 |
| iHOP | AT1G76890 |
| wikigenes | AT1G76890 |



