 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Spipo11G0028200 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Araceae; Lemnoideae; Spirodela
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 236aa MW: 25403.1 Da PI: 4.9901 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.2 | 9.6e-19 | 50 | 100 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+ y+GVr+++ g+Wv+e+r+p +k+ r++lg+f tae+Aa+a++ a+ +l+g
Spipo11G0028200 50 PVYRGVRRRS-AGKWVCEVREP---NKKSRIWLGTFPTAEMAARAHDVAAMALRG 100
68*****888.8******9998...347*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 5.1E-14 | 50 | 100 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 3.33E-21 | 51 | 110 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 2.2E-32 | 51 | 110 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.853 | 51 | 108 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.2E-30 | 51 | 114 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.8E-9 | 52 | 63 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 3.27E-28 | 52 | 110 | No hit | No description |
| PRINTS | PR00367 | 3.8E-9 | 74 | 90 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009631 | Biological Process | cold acclimation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 236 aa Download sequence Send to blast |
MSGYSTEMDS PSGGTAQSGG SDEEVYYATV SSAPPKRRAG RTKFRETRHP VYRGVRRRSA 60 GKWVCEVREP NKKSRIWLGT FPTAEMAARA HDVAAMALRG RSACLNFADS AWRLPVLPGS 120 ATAREIQQAA AKAVQTFRPE GTGADEGGAV GTPSSDTADT ADTADTADTT PPPAEAPASE 180 LEVGYLDLDF GLEDYLANMR MLSPLHPIQM HHHWGEGIWD DVEAESDVSL WSYSI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 1e-14 | 49 | 107 | 12 | 71 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
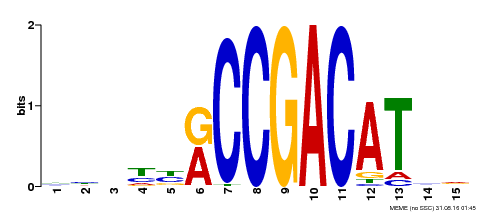
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00452 | DAP | Transfer from AT4G25470 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010255545.1 | 1e-70 | PREDICTED: dehydration-responsive element-binding protein 1B-like | ||||
| TrEMBL | Q0ZDG8 | 3e-70 | Q0ZDG8_9LILI; CBF-like transcription factor 1 | ||||
| STRING | XP_010255545.1 | 4e-70 | (Nelumbo nucifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP196 | 36 | 261 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G25470.1 | 2e-47 | C-repeat/DRE binding factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Spipo11G0028200 |



