 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_108370_teay.t1 | ||||||||
| Common Name | SOVF_108370 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 442aa MW: 48546 Da PI: 6.2943 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.5 | 2.3e-16 | 53 | 97 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE++++++a k++G W++I ++g +t+ q++s+ qk+
Sp_108370_teay.t1 53 RERWTEEEHKKFLEALKLYGRA-WRKIEDHVG-SKTAVQIRSHAQKF 97
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.36E-16 | 47 | 101 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.231 | 48 | 102 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 4.1E-17 | 51 | 100 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.2E-13 | 52 | 100 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.5E-13 | 53 | 96 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-9 | 53 | 93 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.93E-10 | 55 | 98 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 442 aa Download sequence Send to blast |
MPCSSSTMVT QEETSVAQSD SGNRNSSVAG AQTGVGEGQI PKIRKPYTIT KQRERWTEEE 60 HKKFLEALKL YGRAWRKIED HVGSKTAVQI RSHAQKFFSK VVRETNNSNA NSVADPIEIP 120 PPRPKRKPSH PYPRKLVLPS RKDMPMEQGS RSVSPNSSLS EQENQSPTSV LSAAASDMLA 180 SADSAAPKSS PSPVSSGNFG DQVAKLQSED NAPQGEVQES KSGSTVPASV KLELFPVKES 240 VHKEESACTK VLKLFGKDVV VPGNPESFTS LSKSPSLDMG EGVLQAGPED FQAADNSVRI 300 VGNAQDHMKF YYVDKNANQV EKPTPLPWWV FFRGMSVPPF VETQEKETQR EGSSGESNSD 360 SVNGGETQDK NWETETESGQ CGPVSDKEIK FVEDSPSLIE KRSSVDKCRK GFVPYKRCLS 420 ERDMESTIGS EDRESQRVRL CL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
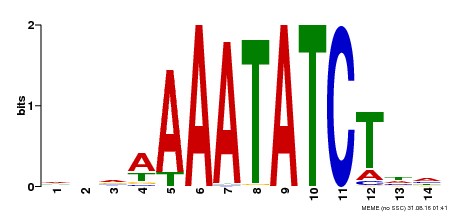
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021856034.1 | 0.0 | protein REVEILLE 1 isoform X1 | ||||
| TrEMBL | A0A0K9R483 | 0.0 | A0A0K9R483_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010675239.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G17300.1 | 9e-43 | MYB_related family protein | ||||



