 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_054560_oaqg.t1 | ||||||||
| Common Name | SOVF_054560 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 293aa MW: 33447.4 Da PI: 4.6852 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 63.1 | 4.1e-20 | 69 | 122 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k++++t+eq+ Le++F +++++ +e++ eLAkklgL+ rqV vWFqNrRa+ k
Sp_054560_oaqg.t1 69 KKRRLTPEQVVLLEKCFDEEKKVEQERKSELAKKLGLQPRQVAVWFQNRRARSK 122
5678************************************************99 PP
| |||||||
| 2 | HD-ZIP_I/II | 107.8 | 8.2e-35 | 68 | 159 | 1 | 92 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelk 92
ekkrrl+ eqv lLE++F+ee+k+e+erK+ela++LglqprqvavWFqnrRAR k+kqlE+d +aLk+++ +l +e ++ ke + L+++++
Sp_054560_oaqg.t1 68 EKKRRLTPEQVVLLEKCFDEEKKVEQERKSELAKKLGLQPRQVAVWFQNRRARSKSKQLERDLDALKASHHTLLSEYQSRLKERDLLKSKVD 159
69************************************************************************999998888888888775 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.27E-18 | 60 | 126 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.572 | 64 | 124 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.5E-18 | 67 | 128 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 8.33E-17 | 69 | 125 | No hit | No description |
| Pfam | PF00046 | 2.1E-17 | 69 | 122 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-19 | 70 | 131 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.1E-6 | 95 | 104 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 99 | 122 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.1E-6 | 104 | 120 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 6.8E-9 | 124 | 166 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001558 | Biological Process | regulation of cell growth | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 293 aa Download sequence Send to blast |
MERDLDSGSL FFDSLANRTN MLFLGNTNLI FPGIRTTMIN MDKKRPFLNL PDEMINDDGY 60 YFDEQLPEKK RRLTPEQVVL LEKCFDEEKK VEQERKSELA KKLGLQPRQV AVWFQNRRAR 120 SKSKQLERDL DALKASHHTL LSEYQSRLKE RDLLKSKVDS LKEKLQSIKG EIEGPITSPI 180 ESLQVDGPTI QAQQQLSKIE DRSSSGSGDS AVVNEENNPQ NMIDSGTSYF DCHPPSPYMV 240 AVQSEEDDIS IDGRTNFSNI ITVAATVAND SDHHQLHEEE EEVGQGFGWW VWA |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 116 | 124 | RRARSKSKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
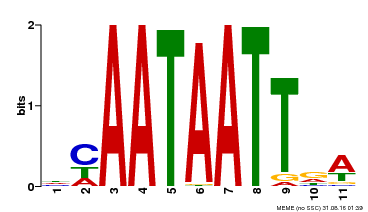
| Motif ID | Method | Source | Motif file |
| MP00323 | DAP | Transfer from AT3G01470 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021866780.1 | 0.0 | homeobox-leucine zipper protein HAT5 | ||||
| TrEMBL | A0A0K9RKV6 | 0.0 | A0A0K9RKV6_SPIOL; Uncharacterized protein | ||||
| STRING | VIT_14s0066g01440.t01 | 3e-80 | (Vitis vinifera) | ||||
| STRING | EOY02359 | 4e-80 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01470.1 | 2e-60 | homeobox 1 | ||||



