 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_004370_hrut.t1 | ||||||||
| Common Name | SOVF_004370 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 381aa MW: 43371 Da PI: 4.7146 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.9 | 4e-19 | 58 | 111 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ +q+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
Sp_004370_hrut.t1 58 KKRRLSVDQVKALERNFEVENKLEPERKVQLAQELGLKPRQVAVWFQNRRARWK 111
566899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 128.8 | 2.3e-41 | 57 | 149 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelk 92
ekkrrls +qvk+LE++Fe e+kLeperKv+la+eLgl+prqvavWFqnrRAR+ktkqlE+dy +Lk+++d+lk++ e+L++++++L +e+k
Sp_004370_hrut.t1 57 EKKRRLSVDQVKALERNFEVENKLEPERKVQLAQELGLKPRQVAVWFQNRRARWKTKQLERDYGVLKSSFDNLKQNFESLQHDKDALLKEIK 148
69**************************************************************************************9998 PP
HD-ZIP_I/II 93 e 93
e
Sp_004370_hrut.t1 149 E 149
7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.3E-19 | 38 | 115 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.281 | 53 | 113 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.1E-17 | 56 | 117 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 2.1E-16 | 58 | 111 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 7.06E-15 | 58 | 114 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 9.4E-20 | 60 | 120 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 8.3E-6 | 84 | 93 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 88 | 111 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 8.3E-6 | 93 | 109 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 3.4E-17 | 113 | 154 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 381 aa Download sequence Send to blast |
MKRTRSSDSL GALISIQEEH SPRNNNNIYS NRDYQSMMLE SLVDEEGCIE EIGHIPEKKR 60 RLSVDQVKAL ERNFEVENKL EPERKVQLAQ ELGLKPRQVA VWFQNRRARW KTKQLERDYG 120 VLKSSFDNLK QNFESLQHDK DALLKEIKEL KSKLGEDRVS KEEEALLFES ENINMNDDDE 180 DDIEPITPPA ESLSGGGGGG GDGGGGSENA KLLKVQNMFA KFKDGGPSDS DSSAILNDDN 240 NNSNSPNASN SSSEPIFQNH QQQQQQQQQQ HHQQQQQLLM SPSSSSFRFN SSSPSPPSIN 300 CFDFSRLDEM RTTQNHNNNN NNNNNNNNNN SNNNSNNSNN VYHPQFVKIE EHNFFSGEEA 360 CNFFSEEQAP SLHWYCPDQW T |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 105 | 113 | RRARWKTKQ |
| 2 | 193 | 206 | SGGGGGGGDGGGGS |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00271 | DAP | Transfer from AT2G22430 | Download |
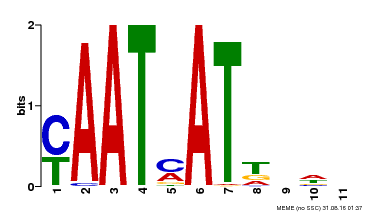
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021852205.1 | 0.0 | homeobox-leucine zipper protein ATHB-16-like | ||||
| TrEMBL | A0A0K9S1U2 | 0.0 | A0A0K9S1U2_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010681015.1 | 1e-135 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22430.1 | 2e-51 | homeobox protein 6 | ||||



