 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sme2.5_01013.1_g00005.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 403aa MW: 46036.2 Da PI: 4.963 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 112.1 | 3.8e-35 | 14 | 105 | 2 | 102 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFk 87
F+ k+ye+++d++++ ++sws+n++sf+v+++ +fa+++Lp+yFkh+nf+SF+RQLn+YgFkk++ e+ weF+++ F
Sme2.5_01013.1_g00005.1 14 FIAKIYEMVDDPSTDPIVSWSSNNKSFIVHNPPDFARDLLPRYFKHNNFSSFIRQLNTYGFKKIHPEQ---------WEFANEDFL 90
9***************************************************************9998.........********* PP
XXXXXXXXXXXXXXX CS
HSF_DNA-bind 88 kgkkellekikrkks 102
+g+ +ll++i r+k
Sme2.5_01013.1_g00005.1 91 RGQPHLLKNIYRRKP 105
************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 8.0E-38 | 8 | 98 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 4.2E-59 | 10 | 103 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| SuperFamily | SSF46785 | 1.9E-35 | 11 | 103 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| PRINTS | PR00056 | 1.0E-19 | 14 | 37 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 2.2E-31 | 14 | 103 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.0E-19 | 52 | 64 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 53 | 77 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.0E-19 | 65 | 77 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 403 aa Download sequence Send to blast |
MDEAPCSTNV LPPFIAKIYE MVDDPSTDPI VSWSSNNKSF IVHNPPDFAR DLLPRYFKHN 60 NFSSFIRQLN TYGFKKIHPE QWEFANEDFL RGQPHLLKNI YRRKPVHSHS AQNLHSVSSS 120 ALTESERQGY KEDIEKLKHE NELLHLELHR HKQDHQELET QLQVLTKRVQ QVKDRQMNVL 180 STLARTINKP GLALSLMPQL EMNERKRRLP GNSLRYNDTG LEDNQVSSSE DLTRENMDPT 240 SLLTLNKEVL DQLESSLTFW EYTLCDIDLA EMRRSSSMDL DESISCADSP AISYPQLTVD 300 VGSKVSDIDV NSEPNGNATP DVTPTDNRVG TASNNVPTGV NDVFWEQFLT ENPGSTDVKP 360 EREDTESKIN ESKTVENGKF WWNRKTVNSL TEQLGHLTPA EGM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 7e-25 | 11 | 103 | 26 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 7e-25 | 11 | 103 | 26 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 7e-25 | 11 | 103 | 26 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 203 | 208 | ERKRRL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
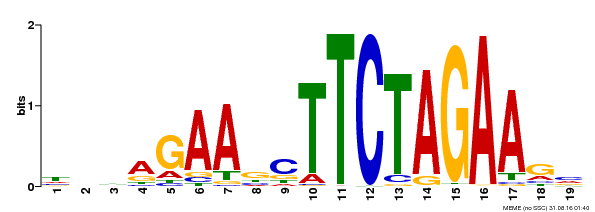
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT014619 | 0.0 | BT014619.1 Lycopersicon esculentum clone 134122F, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015168157.1 | 0.0 | PREDICTED: heat stress transcription factor A-4a-like | ||||
| Refseq | XP_015168158.1 | 0.0 | PREDICTED: heat stress transcription factor A-4a-like | ||||
| TrEMBL | M1CNW0 | 0.0 | M1CNW0_SOLTU; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400071487 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3448 | 23 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18880.1 | 4e-97 | heat shock transcription factor A4A | ||||



