 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Solyc10g005080.2.1 | ||||||||
| Common Name | LOC101261662 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum; Lycopersicon
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 763aa MW: 83707.9 Da PI: 6.4 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50.5 | 4.6e-16 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE+ ++++a k++G W +I +++g ++t+ q++s+ qk+
Solyc10g005080.2.1 24 RERWTEEEHNRFLEALKLYGRA-WQRIEEHIG-TKTAVQIRSHAQKF 68
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 8.22E-17 | 18 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.173 | 19 | 73 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 7.0E-17 | 22 | 71 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.1E-12 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-8 | 24 | 64 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 3.2E-13 | 24 | 67 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.48E-9 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010243 | Biological Process | response to organonitrogen compound | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043496 | Biological Process | regulation of protein homodimerization activity | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 763 aa Download sequence Send to blast |
MDPYSSGEEL VVKTRKPYTI TKQRERWTEE EHNRFLEALK LYGRAWQRIE EHIGTKTAVQ 60 IRSHAQKFFT KLEKEALTKG VPTSQALDIE IPPPRPKRKP SNPYPRKTSA AGHSSQVGAK 120 DGKYSTTFSS ICEERNLFDL EKEPITEKPG GNEKLGNVKE TQNKKNCSQG LTKEGASAAS 180 MSSGKSLQAH VAPTDVCAFS ESVSVTKGVV NNDNANKSFL IVESKEHQQS EILDIRQSFQ 240 GNSSCNTFDG GKSCQSSEKL AQGEKKHPSF QPNHLGEFSR NDMQVLHNYP RHVPVHILDG 300 TNGSQIAPDM FNHESTSQQI NGVPGLPNLY SNPASSTTSE HHSNAPQSSI HQSFSCFHPI 360 FTPIRDPDDY RSFFQLSSTF SSLIVSALLQ NPAAHVAASF AASFWPYANM ERPTDSPTDN 420 TASQINSAPS MAAIAAATVA AATAWWAAHG LLPLCSQFQS SFTCVPTSAT SMQVDACQPR 480 VDKNEGREGT HDSPHVQEPV PECSEALQEQ QSGSKLPPSL SSESEESEGR KLKTGLTATD 540 TEQGAAVTKI NEPNAEKGGK QVDRSSCGSN TPSSSEIETD ALEKDEKGKE EPQESNINLL 600 AGEAANRRYR NFISPTESWK EVSEEGRIAF QALFTREVLP QSFSPSLDLK NKGKIILEKL 660 KQKPDEKVQC GPQLDLNDMA SNICSSHQTM EDNVLLIGNK EDVETCLPMI ELGQVRLKAR 720 RTGFKPYKRC SLEANDSRVT SSNCQDEEKS SKRLRLEGEA ST* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Les.4923 | 0.0 | flower| fruit| leaf| root | ||||
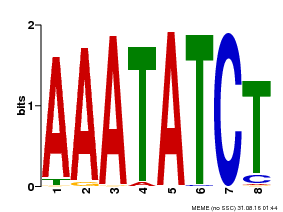
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00103 | PBM | Transfer from AT2G46830 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Solyc10g005080.2.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT012912 | 0.0 | Lycopersicon esculentum clone 114030R, mRNA sequence | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004248416.1 | 0.0 | protein LHY isoform X2 | ||||
| Refseq | XP_010327245.1 | 0.0 | protein LHY isoform X2 | ||||
| Refseq | XP_019071645.1 | 0.0 | protein LHY isoform X2 | ||||
| Refseq | XP_025883775.1 | 0.0 | protein LHY isoform X2 | ||||
| TrEMBL | A0A3Q7IAI7 | 0.0 | A0A3Q7IAI7_SOLLC; Uncharacterized protein | ||||
| STRING | Solyc10g005080.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA9710 | 13 | 15 | Representative plant | OGRP9903 | 5 | 10 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01060.3 | 2e-49 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Solyc10g005080.2.1 |
| Entrez Gene | 101261662 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



