 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Solyc03g111590.2.1 | ||||||||
| Common Name | LOC101257458 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum; Lycopersicon
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1253aa MW: 139814 Da PI: 9.1879 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.3 | 0.0011 | 1136 | 1161 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y C C++sFs+k +L+ H ++ +
Solyc03g111590.2.1 1136 YHCDleGCSMSFSSKQELTLHKKNvC 1161
889999***************99866 PP
| |||||||
| 2 | zf-C2H2 | 11 | 0.0014 | 1160 | 1183 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirtH 23
+Cp C+k F ++ +L++H r+H
Solyc03g111590.2.1 1160 VCPveGCKKKFFSHKYLVQHRRVH 1183
69999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 11.1 | 0.0013 | 1219 | 1245 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y C+ Cg++F+ s++ rH r+ H
Solyc03g111590.2.1 1219 YACSeiGCGQTFRFVSDFSRHKRKtgH 1245
889999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 3.6E-16 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.991 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 2.1E-15 | 21 | 54 | IPR003349 | JmjN domain |
| SMART | SM00558 | 5.8E-49 | 184 | 353 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 32.372 | 187 | 353 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 8.79E-25 | 199 | 351 | No hit | No description |
| Pfam | PF02373 | 1.9E-36 | 217 | 336 | IPR003347 | JmjC domain |
| SMART | SM00355 | 8.6 | 1136 | 1158 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.572 | 1159 | 1188 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.012 | 1159 | 1183 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 4.3E-5 | 1159 | 1187 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1161 | 1183 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.39E-9 | 1175 | 1217 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.0E-8 | 1188 | 1213 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.741 | 1189 | 1218 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 1189 | 1213 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1191 | 1213 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.56E-7 | 1207 | 1241 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.7E-8 | 1214 | 1241 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.074 | 1219 | 1250 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.48 | 1219 | 1245 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1221 | 1245 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1253 aa Download sequence Send to blast |
MAEASGNIEV FSWLKTLPVA PEYHPTLEEF QDPIAYIFKI EKEASKYGIC KIVPPVPAPP 60 KKTALANLNR SLSARAGSNG PTFTTRQQQI GFCPRKHRPV KKPVWQSGET YTVQQFQVKA 120 KAFEKNYLRK NSKRALTPLE VETLYWKATV DKPFSVEYAN DMPGSAFAPK KASLAAGGIG 180 EVSTLADTEW NMRGVSRSKG SLLKFMKEEI PGVTSPMVYL AMMFSWFAWH VEDHDLHSLN 240 YLHMGSGKTW YGVPRDAAVA FEEVIRVQGY AGETNPLVTF ATLGEKTTVM SPEVLLSAGI 300 PCCRLVQNAG EFVVTFPRAY HSGFSHGFNC GEASNIATPE WLRVAKDAAI RRASINCPPM 360 VSHFQLLYDL ALSLCSRVPK NIRIEPRSSR LKDKKKSEGD MLVKELFVED LNANNYLLHI 420 LGEGSPVVLL PQNSPGISIC SNLVAGSQSK VNSRLFPSSS NSDHEVKSKK DSAYDDRKLG 480 RKQGMKQYAG ISLEKGKYSS WHTGNSLPDS GRKDDAQSSP ETEKVNLDAA RGMTYKCDTL 540 SEQGLFSCAT CGILCYTCVA IIRPTEAAAR HLMSSDYSDF NGWTGSVSGI TATGRDPNAA 600 ESDSSSGRFV KRAPALIDDP VESSDRIQKL NNGSVEELSR TNTRKETSSL GLLALAYANS 660 SDSDEDEIEV DIPVEACESR HTESEDEVFL RVIDPYGNHR QKRAVSQGRN CQKFDNSVQL 720 ENESYPSGES NTLFGRSSHQ PRSHQVPAKC ISNIREIAQN NAVAPFDNAR MQFTSTSDED 780 SFRIHVFCLQ HAVQVEEQLR RIGGAHISLL CHPDYPKLEA QAKQVAEELG SDHFWREISF 840 REASKEDEEM IQSALEIEEA IHGNGDWTVK LDINLFYSAN LSRSPLYSKQ MPYNFIIYNA 900 FGRDSPDNTP EKSEYTGRGL GKQRRAIVAG KWCGKVWMSS QVHPLLAERT IDEEQEQNKS 960 ISALIKIEVK SERPRERTPT SKTVATTCKT GKKRSSTAAS RNASNAQLII ADDHDDSLLS 1020 SILQQHRRKT NLRSKRIKYE TPEPQKDVDK KKIFGSLIDD DPDGGPSTRL RKRIPKPSNE 1080 SPAKSVKAKP APTKQHESKK GPKVKLPFAN SIAKKEPVTK GPRSNIGKRM REEGEYHCDL 1140 EGCSMSFSSK QELTLHKKNV CPVEGCKKKF FSHKYLVQHR RVHMDDRPLK CPWKGCKMTF 1200 KWAWARTEHI RVHTGARPYA CSEIGCGQTF RFVSDFSRHK RKTGHISKKG RS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 3e-73 | 13 | 386 | 8 | 366 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 3e-73 | 13 | 386 | 8 | 366 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in the shoot apical meristem and primary and secondary root tips, and lower expression in cotyledons, leaves and root axis along vascular tissues. Detected in inflorescences, stems and siliques. {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18713399}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
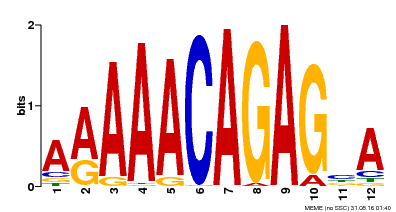
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Solyc03g111590.2.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC238563 | 0.0 | Solanum lycopersicum chromosome 3 clone C03HBa0049N20, complete sequence | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004236313.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A3Q7FPK7 | 0.0 | A0A3Q7FPK7_SOLLC; Uncharacterized protein | ||||
| STRING | Solyc03g111590.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4988 | 23 | 31 | Representative plant | OGRP4401 | 12 | 17 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Solyc03g111590.2.1 |
| Entrez Gene | 101257458 |



